Welcome to the OLI version of Modern Biology. This introductory course is called "Modern
Biology" because it is focused on topics at the forefront of experimentation in the
fields of cellular biology, molecular biology, biochemistry, and genetics. It is carefully
planned to provide the background students will need for advanced biology classes.
Students from other disciplines may also find this course useful as it explains many of
the concepts and techniques currently discussed in the popular press and applied in
other contexts.
Fluorescent image of a living mouse cell
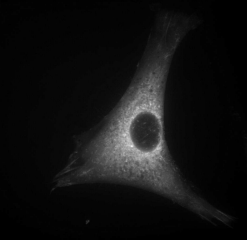 This is a mouse tissue culture cell showing the cytoskeletal structure. The
large clearing in the center is the nucleus. Image courtesy of Jon Jarvik,
Department of Biological Sciences, Carnegie Mellon University.
This is a mouse tissue culture cell showing the cytoskeletal structure. The
large clearing in the center is the nucleus. Image courtesy of Jon Jarvik,
Department of Biological Sciences, Carnegie Mellon University.
An image of a living mouse cell in culture is shown above. As the cell adheres to the
bottom of the culture dish it spreads out exploring the local environment giving it
the angular shape. The cell can move, utilize energy, and divide to produce new cells.
During this course you will explore the fundamentals of how a cell is able to carry out
each of these processes. Modern biology is about the molecular events that occur inside
a cell: the making of proteins, the building of cellular structures, and the interaction
of a cell with its environment.
This Modern Biology course is built around several key concepts that provide unifying
explanations for how and why structures are formed and processes occur in a biological
system. Because it is not possible to cover the breadth of modern molecular biology in
one semester, an understanding of these key concepts will provide a basis for extension
of your knowledge to biological systems beyond the specific topics covered in this
course. One of the major goals of the course therefore is for you to not only learn the
fundamentals of the concepts but also to recognize how they can be applied in other
contexts. Several key concepts include:
- Bioselectivity
- Energy
- Equilibrium
- Ionic State
- Regulation
- Solubility
- Correlation of Structure and Function
The course is organized into units covering the areas of basic biochemistry, cell
biology, and molecular and cellular function. The first unit introduces the basic
chemistry of a cell. All other units will rely heavily on the concepts and background
introduced in this unit. You are encouraged to master this material before proceeding to
the other units.
Before you begin you may want to read the General Instructions about the course.
To put the course in perspective, we begin by exploring the cell
and the components of the cell called organelles. The focus of this
course is to understand the components of the cell, how they interact
with each other, how they are created, destroyed and how they regulate
transport, growth and division of the cell. We will examine the
controlled chemical environment a cell maintains and what restrictions
this places on the types of chemical reactions it can perform. This
background is vital
to understanding key processes such as how a cell releases energy
from glucose, makes
and folds proteins, and goes through growth and cell division.
Above is a caricature of a eukaryotic cell. Many of the cell components are hyper links that will
provide you with an image showing these same structures in a living cell. This
illustration highlights one of the goals of the course which is to expand your view of
biology by bridging from classic simple illustrations to images generated from actual
data. In addition, you will develop an understanding of the fundamental processes used
in this imaging method.
Practice
This exercise uses the Cell to let you test your knowledge
of the functions of the organelles.
Biological systems use only a small subset of the elements (approximately 10 %) found in
the periodic table. The chemical reactions that take place in cells represent only a
small subset of all possible reactions. Before we begin an in-depth study of other
aspects of molecular and cellular biology, one needs to understand the restrictions the
cellular environment places on the possible chemical reactions and the resulting
structures. By learning about the characteristics of the subset atoms, and a limited set
of functional groups found in biological molecules, you will be able to identify, and
predict many of the reactions that can take place, understand and predict the physical
properties of the molecules made by these reactions, and develop an understanding of why
a process occurs as it does in a cell.
Roll over each axis for details on the constraints biological systems make
on that axis. The blue cube represents the chemical universe as defined by
temperature (in Kelvin) on one axis, pH on another, and the elements on the third.
The small orange interior cube represents the part of chemistry that occurs in
cells. 310K is average human body temperature (37˚C or 98.6˚F).
We start with an understanding that the cellular environment is essentially aqueous
(water) based, and thus, we will begin with a discussion of the chemistry of water as it
relates to bonding, pH and temperature control. The first two concepts (bonding and pH)
dictate why molecules are soluble in the cell, in what part of the cell they are
soluble, what charge molecules will carry and how that charge is controlled by pH.
In the next few pages, we discuss each of the 10 learning objectives described in the
objective link (check mark icon at the top and bottom of each page). Each subsequent
page will list the specific objectives relevant to that page. Many of the simulations
and demonstrations, used in this module, are referenced in the Glossary Module.
Atoms of Life
The key biologically relevant elements are hydrogen (H), carbon
(C), nitrogen (N), oxygen (O), phosphorous
(P), and sulfur (S). These elements represent over 95% of the
mass of a cell. Carbon is a major component of nearly all biological molecules.
Some elements are found in relatively small amounts and are called “trace
elements.” The examples include sodium (Na), potassium (K), chlorine (Cl),
manganese (Mn), Zinc (Zn). Throughout the course you will see how atoms of these
elements are very important to the functioning of a cell. Living organisms get
the required elements from outside and constantly rearrange these elements to
build their own molecules. Thus, understanding behavior and structure of
elements is important for understanding life.
Elements are characterized by their atomic structure. While the subatomic
structure of the atom is a major topic of interest in chemistry, physics and
biophysics, the basic structure described below provides sufficient information
for the construction of molecules in the context of this course. Atoms are made
up of subatomic particles: protons, neutrons, and electrons. Protons and
neutrons are at the center of the atom and have a mass of 1 atomic mass unit
(a.m.u) each. Each proton has a positive charge (+1), while neutrons are neutral
(they carry no charge). Each electron has a negative charge (-1) and zero mass.
Two atoms that differ by the number of neutrons are called isotopes of the same
element (e.g. radioactive isotope of iodine is used for cancer treatment).
These elements represent over 95% of the mass of a cell. Carbon is a major
component of nearly all biological molecules. Elements are characterized by
their atomic structure. While the subatomic structure of the atom is a major
topic of interest in chemistry, physics and biophysics, the basic structure
described below provides sufficient information for the construction of
molecules in the context of this course. Atomic mass, the sum of the number of
protons and neutrons in the atomic structure, is a particularly useful measure
of each element. By summing the atomic mass of all the atoms in a molecule, one
can estimate the molecular mass of the molecule, which is then expressed in
atomic mass units or Daltons. The masses of the six atoms of the elements listed
above are given in the following Atomic Properties TABLE. The masses
can be found in the upper right hand corner of the box for each element in the
periodic table.
| Atomic Properties of the Major Biological Atoms |
|---|
| Atom |
Mass |
| H |
1 |
| C |
12 |
| N |
14 |
| O |
16 |
| P |
31 |
| S |
32 |
|
Atoms Form Molecules
One characteristic of the atoms of the major elements is that they are
able to form molecules through formation of covalent bonds with other
atoms.
- Covalent bonds
-
(Definition)
Covalent bonds represent the sharing of the electrons
(negatively charged subatomic particles between atoms.) The
number of covalent bonds that can form is dictated by the
number of unpaired electrons in the outer valence shell of
the atom.
Each atom in a molecule will complete its outer shell of electrons,
which is 2 for hydrogen, and 8 for second row elements (e.g. C, N and
O). The valence shells for each of the biologically relevant elements
are highlighted in the periodic table below. The relationship between
the number of unpaired electrons in the valence shell and the number of
possible covalent bonds an element can form is given in the Atomic
Properties TABLE above.
Only the valence shells are shown. The six shaded elements have
unpaired electrons and readily form covalent bonds.
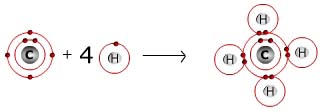
Molecules are made up of atoms covalently bound to each other. For example, a
molecule of methane is a carbon atom covalently bonded to four hydrogen atoms;
and water is composed of an oxygen atom covalently bonded to two hydrogen atoms.
Molecules can also be complex containing many atoms covalently bonded to each
other, for example, cholesterol. Note that cholesterol also contains bonds that
involve the sharing of more than one valence electron between two atoms creating
a double bond indicated by C=C. In this case, each carbon still only
participates in making four bonds. This will be discussed later in the section
on bonding.
Electronegativity
Another property of the atoms is electronegativity.
- Electronegativity
-
(Definition)
The tendency of an atom to attract electrons to itself.
Electronegativity increases as one moves from left to right, across
the periodic chart. Because the electronegative atoms have the
potential to attract electrons, i.e., the electrons spend more time
on the electronegative atom, a molecule containing an
electronegative atom will have partial negative charge associated
with that atom (as indicated by δ-). The the
bonding partner in the covalent bond becomes partially positively
charged (as indicated by δ+ ).
The arrow shows the tendency of atoms of the various elements to lose,
keep, or gain an electron.
As you can see in the table, oxygen, nitrogen, and
sulfur are electronegative atoms and represent the major electronegative atoms
in biological molecules. Phosphorous, an important component in nucleic acids
(e.g. DNA) is also electronegative. Water is a good example of how the very
electronegative atom of oxygen influences the electrons shared with hydrogen.
Oxygen has six valence electrons; two sets of paired electrons and two sets of
single bonding electrons. The shared electron pair of each oxygen-hydrogen
covalent bond, illustrated below, spends more time associated with the oxygen
atom than the hydrogen atom. This results in the oxygen being partially
negative, and the hydrogen partially positive, as indicated in the figure below
by δ- and δ+. These are then described as a
polar covalent bonds. The presence of polar covalent bonds in water
and in other molecules containing electronegative atoms puts these molecules in
a family of molecules referred to as being polar. We will return to
polar bonds in water and their consequences in the "Importance of Water"
section. Oxygen is not the only electronegative element that forms polar bonds,
as long as the electronegativities of the two atoms differ, the bond will have
some degree of polar character. For example, the P-O bond in DNA is polar, with
the phosphorous having a partial positive charge since it is less
electronegative than oxygen.
Because oxygen is strongly electronegative, it draws the electrons, e-,
it shares with the hydrogen atoms, to itself creating a charge imbalance as
indicated by the bold arrow. The oxygen is slightly negative and the
hydrogen is slightly positive. This imparts a polar characteristic to the
oxygen-hydrogen bond.
The Ionic State
- Ionic Bond
-
(Definition)
An
ion is an atom or a molecule that carries a charge. Negatively charged ions,
called anions, form when a neutral molecule or atom gains one or more electrons.
Positively charged ions, called cations, form when a neutral molecule or atom looses one
or more electrons.
Ionic bonds are interactions between oppositely charged ions. An ionic bond forms
due to an attraction between a positive and a negative ion. No electron sharing occurs in
the ionic bond.
There are trace amounts of many other elements in cells most of which exist as
ions. These include sodium (Na+), chlorine
(Cl-), fluorine (F-), iron (Fe++),
magnesium (Mg++), cobalt (Co++), and manganese
(Mn++). Throughout the course you will see how atoms of these
elements are very important to the functioning of a cell.
As you can see in the periodic table these atoms are found on the extreme right
or the extreme left of the periodic table. For example, chlorine is on the right
of the periodic table and is extremely electronegative and, thus, wants to
acquire an electron, whereas, sodium on the extreme left of the periodic table
wants to give up its unpaired electron. By losing an electron, sodium becomes a
positively charged cation and by gaining an electron, chlorine becomes
a negatively charged anion. These charged atoms are called
ions and form the basis for charge repulsion and attraction in the
non-covalent ionic bond. The affinity of Na+ for Cl- is a
non-covalent, ionic bond (the attraction of opposite charges).
Water and Hydrogen Bonding
Water is the solvent of life on Earth. It has several properties that contribute
to its suitability to support life as we know it. One property derived from the
special properties of oxygen is that water is a polar
molecule. Oxygen is electronegative and draws the electrons that it
shares in the covalent bond with hydrogen towards itself.
The electronegative oxygen (red) draws electrons to it, creating
the partial negative charge on oxygen and partial positive charge on
hydrogen.
Water Molecular Structures
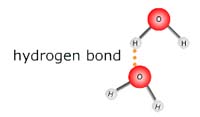
In pure water, the partially negative oxygen of one molecule attracts
the partially positive hydrogens from another water molecules to form a
non-covalent bonding interaction called a hydrogen bond.
- Hydrogen Bonding
-
(Definition)
The attraction of an electronegative atom for a hydrogen that is
covalently bonded to another electronegative atom. This involves the
attraction of a hydrogen with a partial positive charge to an atom with
a partial negative charge. However, only hydrogens covalently bonded to
an electronegative atom can participate in hydrogen bonding.
Adjust the volume on your computer and click the play
button.
Answering questions like the one above is good practice for answering the kinds of
questions that may appear on the exams. Your answers may also shape discussions in
class.
Molecular Bonding
Three major types of chemical bonding have been described thus far: covalent bonding,
and two forms of non-covalent bonding, ionic and hydrogen. These bonds are all important in
the functioning of a cell.
Covalent Bonds
Covalent bonds are the strongest. One atom fully shares one, two or three
electrons with another atom., forming a single, double, or triple bond,
respectively. The bonds can be between the same element (e.g., C-C bonds) or
between different elements (e.g., C-O, C-N, H-O). The nature of the covalent
bond is determined by the number of electrons shared and the nature of the two
elements attached.
Single bonds: Two atoms attached by a single covalent bond have free
rotation about the bond.
Double bonds: Two carbons attached by a double covalent bond can only
undergo 180 degree rotations and the atoms bound to these carbons are
constrained to lie in the same plane as the carbon atom.. Rotation about the
carbon-carbon bond has structural implications for molecules in which they are
found. For example, the type of bond can influence the fluidity of biological
membranes and restrict the folding of proteins.
Triple bonds. Although carbon can form triple bonds, such as in the
compound acetylene (HCCH), triple bonds are not found in biological systems.
Aromatic Compounds: Involve the sharing of electrons between atoms that
form a ring. The shared electrons form a partial (1/2) double bond between each
atom in the ring structure. Aromatic compounds have unique geometrical
properties and absorb ultraviolet light .
Formation of a covalent bond
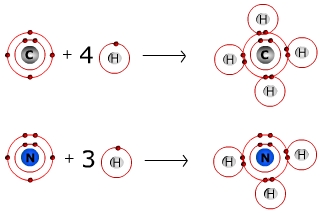 A covalent bond is formed between two hydrogen atoms.
A covalent bond is formed between two hydrogen atoms.
Covalent bonds involving electronegative atoms often result in polar molecules.
As discussed earlier, the bonds between the hydrogen and oxygen atoms of water
are polar, due to the fact that oxygen is electron withdrawing and hydrogen is
willing to give up its lone electron. The double bond between carbon and oxygen
atoms in a carbonyl group is polar, for the same reason that the oxygen attracts
the electrons to itself.
Geometry
The overall shape of molecules depends on the geometry of the bonds that are
formed between atoms. The shape of a molecule can have a large effect on its
biological activity, often small changes in the shape of a molecule will make it
biologically inactive.
We saw with water that the orientation of atoms around the oxygen was tetrahydral
with the angle between the two hydrogen atoms close what would be expected for a
tetrahydral shape. Both carbon and nitrogen also form tetrahydral shapes. In the
case of methane the four hydrogen atoms are at the corners of a tetrahydron. In
ammonia, the three attached hydrogens form the base of a tetrahydron. In the
ammonium ion, the addition of a fourth hydrogen, to form
NH4+, places the fourth hydrogen at the last unoccupied
corner of the tetrahydron.
Carbon and nitrogen can also form planer geometries, where all of
the atoms that
are bound to the carbon (or nitrogen) are in the same plane as the
carbon or
nitrogen. In the case of carbon, a planer geometry is usually
observed if the
carbon is involved in a double bond, while the tetrahydral geometry
is found if
the carbon is forming single bonds. Compare the structure of
methanol to
formaldhyde in the Jmol below. In methanol, carbon forms four
single bonds while
in formaldehyde there is a double bond between the carbon and the
oxygen. The carbon atom in methanol shows tetrahdyral geometry, while
the carbon atom in formaldehdye is planer; the two hydrogens and oxygen
lie in the same plane as the carbon..
Chirality
An important aspect of carbon bonding is the fact that
carbon can covalently bond to four groups and that the bonding geometry of the
carbon atom is tetrahedral. Therefore, if the four groups attached to the carbon
are different, then two unique arrangements of the groups around the carbon atom
are possible and this carbon is said to be an asymmetric center or a
chiral center. The two arrangements of groups about the carbon are
mirror images of each other and these two structures are referred to as
enantiomers. It is impossible to superimpose these two mirror
images, consequently they are distinct molecules.
Because the enantiomers have identical functional groups attached to the chiral
center they have identical physical and chemical properties - except for the
direction they rotate plane polarized light. Experimentally, the enantiomers are
distinguished by the direction of rotation of plane polarized light. The
enantiomer that rotates light to the right is designated as D (dextro) form of
the compound. The other enantiomer will rotate light in the opposite direction,
and is designated as the L (levulo) form of the compound.
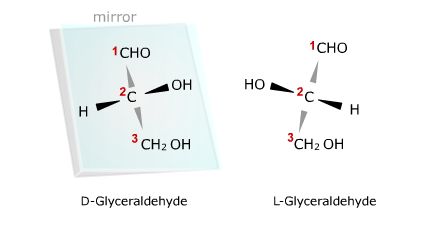
The example shown below is the three carbon carbohydrate, glyceraldehyde. This
compound contains a chiral center at the middle, or second carbon, because that
carbon has four different groups attached to it. The configuration of atoms in
the left-hand structure causes polarized light to be rotated to the right and is
therefore the D form. Its mirror image shown on the right, rotates polarized
light in the opposite direction, and is therefore the L form. Note that in both
compounds the H and OH groups project out of the page towards you, but in the D
form the -OH group is to the right of carbon two while in the L-form it points
to the left.
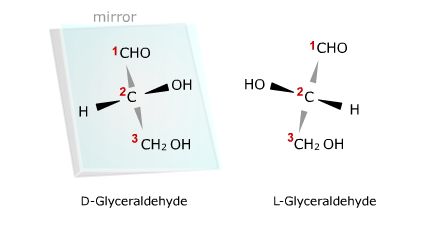 The two arrangements of atoms around a chiral center are mirror images
much as the right hand is a mirror image of the left. The dark black arrows
indicate that atoms are above the page and the gray arrows indicate the
atoms are below the page.
The two arrangements of atoms around a chiral center are mirror images
much as the right hand is a mirror image of the left. The dark black arrows
indicate that atoms are above the page and the gray arrows indicate the
atoms are below the page.
While enantiomers usually undergo chemical reactions in an identical fashion,
biological systems are capable of discriminating between these structurally
different molecules because biological systems themselves contain chiral
centers, such as in amino acids. The ability to discriminate between the
enantiomers is an example of bioselectivity. The chemical world has been
narrowed by the selective use of specific enantiomers (in this case only
D-glyceraldehyde) by biological systems.
- chirality
-
(Definition)
When identical groups attached to a carbon are arranged in multiple
ways such that two of the resulting structures are
non-superimposable, they are mirror images of each other.
Ionic Bonds
Ionic bonds form between oppositely charged atoms. No electron sharing or
transfer occurs. The atoms are attracted to each other due to their opposite
charges. For example, the positive Na ion, and negatively charged Cl ion, are
attracted to each other and form table salt. In an aqueous solution, these ions
are completely dissociated and are defined as strong electrolytes. In water
molecules surround the ions to form polar interactions to satisfy the charges on
the ions. Thus the ions become encapsulated by water spheres, which are called
spheres of hydration. The biological world is very ionic and the
spheres of hydration are important in a cell because they maintain the
separation of the many ions of the cell from each other. The sphere of hydration
must be broken in order for binding to take place with a specific binding
partner.
Previously, water was described as having a high dielectric constant. This
property that is a measure of the polarity of the covalent bond within the
molecule accounts for the separation of ions by polar molecules such as water.
The force of attraction between two oppositely charged ions is inversely
proportional to the dielectric constant. Thus water with a high dielectric
constant decreases the attraction between opposite charges. This is physically
explained by the ability of polar solvents to form ordered hydration layers
around ions.
Hydrogen Bonding
Hydrogen bonding was covered in Water -- Hydrogen Bonding (go there now.) Remember
that hydrogen bonding occurs between partially negatively charged
electronegative atoms, and partially positively charged hydrogen atoms that are
attached to electronegative atoms such as oxygen, nitrogen or sulfur. Hydrogen
bonding is a critical bonding in the cell. It is the principal bonding that
holds the tertiary structure of proteins, carbohydrates and nucleic acids
together and the overall stability of these molecules is due in part to the
cumulative effect of the large number of hydrogen bonds found in the functional
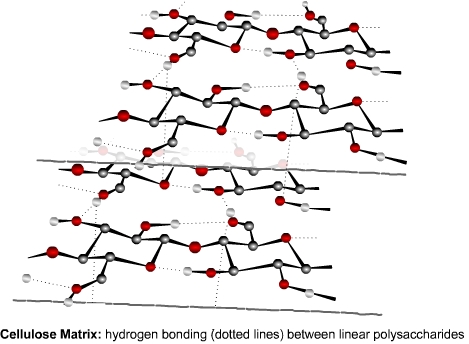
structures. Hydrogen bonds are found in and between a variety of molecules. For
example, the enormous number of hydrogen bonds between strands of cellulose
provide the strength and structure of the plant cell wall.
Hydrophilic Interaction
The nature of polar molecules is that they contain electronegative atoms,
consequently they are capable of hydrogen bonding with aqueous or polar
solvents. Because polar molecules are generally water soluble, they are referred
to as being hydrophilic, or water-loving. The one-carbon alcohol,
methanol, is an example of a polar molecule.
Hydrophobic Interaction
The final type of interaction occurs between neutral, hydrophobic, or
water-fearing, molecules. These molecules do not interact with water and are
characterized by a complete lack of electronegative atoms. In aqueous solutions
the hydrophobic molecules are driven together to the exclusion of water. For
example, shaking a bottle of oil and vinegar (acetic acid in water), such as in
a salad dressing, results in the oil being dispersed as tiny droplets in the
vinegar. As the mixture settles, the oil collects in larger and larger drops
until it only exists as a layer, or phase, above the vinegar.
A similar effect occurs in biological systems. As a protein folds to its final
three-dimensional structure, the hydrophobic parts of the protein are forced
together and away from the aqueous environment of the cell. Similarly,
biological membranes are stabilized by the exclusion of water between layers of
lipids as we will see later.
The hydrophobic effect does not involve direct bonding between the non-polar
molecules, it is an entropy driven process. You may recall that processes that
increase the disorder of a system are more favorable. When a hydrophobic
molecule is truly dissolved in water, the water forms a highly ordered ice-like
shell around the compound. When the hydrophobic molecules contact each during
separation of the aqueous and non-polar phases, the ordered water is released
and become highly disordered. The increase in disorder of the released water
molecules is responsible for the spontaneous assembly of many biological
systems, such as proteins and membranes.
Mixed Non-polar/polar molecules: Of course, there are instances where
even molecules with electronegative atoms will not be water soluble. Computer
algorithms are currently used to predict water solubility based on structure.
For our purposes, we will balance the ratio of polar and non-polar elements in a
structure to estimate the chemical nature of any compound we are going to study.
Amphipathic molecules are molecules that have a distinct non-polar, or
hydrophobic region, and a distinct polar region. These molecules do not form
true solution is water. Rather, the non-polar parts are forced together into a
non-polar aggregate, leaving the polar part of the molecule to interact with the
aqueous phase. Detergents and long-chain carboxylic acids are examples of
amphipathic molecules.
van der Waals Interactions
An important force in biochemistry is due to van der Waals interactions.
This interaction occurs between any two surfaces that are in contact. The force
is actually an electrostatic one that occurs as a result of a momentary
fluctuation in the charge on one surface.. This charge causes the other surface
to momentarily assume the opposite charge, leading to a net attractive force. If
one of the surfaces has a permanent dipole, due to the presence of
electronegative or electropositive atoms, then the attraction is stronger. The
strength of van der Waals forces depend on the contact surface area; the larger
the area the larger the interaction. At the molecular level, van der Waals
interactions can contribute 10s of kJ/mol of energy. At the macroscopic level
van der Waals forces can become quite large. For example, the common gecko
generates sufficient van der Waals forces due to the large surface area of its
foot pads to walk on the ceiling!
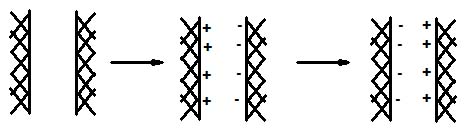 Origin of van der Waals effects. Two neutral surfaces (left) have no
net attraction. One surface becomes charged for a short period of time. The
charge on one surface generates a charge of the opposite sign on the other
surface, leading to an attractive force between the two surfaces. A short
time later the charges reverse in sign, again generating an attractive force
between the two surfaces.
Origin of van der Waals effects. Two neutral surfaces (left) have no
net attraction. One surface becomes charged for a short period of time. The
charge on one surface generates a charge of the opposite sign on the other
surface, leading to an attractive force between the two surfaces. A short
time later the charges reverse in sign, again generating an attractive force
between the two surfaces.
Energy Associated with the Bonds
Each of the bond types represents a measurable amount of energy. To break a
bond, the equivalent amount of energy must be expended. In metabolism, bonds are
broken in molecules, such as glucose, to "release" the energy. The cell utilizes
this energy to drive other energy consuming reactions. The covalent bond has the
most energy associated with it, on average approximately 100 kilocalories/mole
(kcal/mol). The non-covalent bonds, ionic and hydrogen, and hydrophobic
interactions, have approximately 5 kcals/mol associated with each of them.
It should be noted here that throughout the presentation of this course
approximations will be used for certain values so that estimations can be made
as we move to more complex systems. It is to be acknowledged that very precise
values for each of the measurements are not available.
Thus the non-covalent bonds that have been introduced have approximately 20
times less energy associated with them and, thus, are more easily broken
individually. However, hydrogen bonds generally form extensive networks, and the
total energy associated with the network is the sum of the individual
interactions. As anyone who has done a "belly buster" knows, breaking a large
surface area of water is extremely difficult (and painful!).
| Energy Associated with the Different Bonds |
|---|
| Bond |
Energy, kcal/mol |
| Covalent |
100 |
| Ionic |
5 |
| Hydrogen |
5 |
| Hydrophobic interactions |
5 |
| van der Waals |
5 (depends on surface area) |
|
When NaCl dissolves in water, each atom becomes surrounded by at least 20 water
molecules. As NaCl there is 5 kcal/mol of energy associated with the ionic
attraction of the cation and anion, but when a Na ion is surrounded by 20 water
molecules, there is 100 kcal/mol of energy associated with just the Na ion.
Thus, NaCl in an aqueous solution is energetically more favored than NaCl as the
ionically bonded molecule due to the resulting hydrated state. You will explore
what happens to molecules that only partially dissociate, or weak electrolytes,
in water in the next module.
Biological systems use only a small part of the total chemical repertoire. This is due
to the number of chemical reactions that can occur under physiological conditions, and
to the small number of chemical or functional groups compatible in biological systems.
The physical properties of these groups define how molecules behave in biological
systems at physiological pH and temperature. By learning the properties of these groups,
you will be able to predict how various molecules will function, as well as which
functional groups can be converted to others during metabolism
The functional groups fall into three broad categories: non-polar, polar neutral, and
polar charged. The molecules in each group all have common properties.
Functional Group Tutorial
If you need a reminder of chemical structures or how to view molecules in 3-D,
use the link below.
Practice
This exercise uses the Functional Group Glossary to let you test your knowledge
of the properties of the functional groups.
In the previous module you learned about a limited set of functional groups on molecules found in
biological systems and some of their properties. Specifically you learned there were only three
types of functional groups: non-polar, polar neutral, and polar charged. Within these types there
are eight that are particularly relevant to biological systems out of the hundreds of known
organic functional groups.
Of the three types, the polar charged groups, amino, carboxyl, and phosphate, undergo the
greatest change depending on the environment in which they are found. They are weak electrolytes
and behave as bases (amino groups) or acids (carboxyl and phosphate groups) and as such have
the potential to exist as ions. In this module you will explore the properties of weak acids
and bases including their dissociation equilibrium and their ability to act as pH buffers. The
cell exploits these properties to control intracellular pH, the ionic state of molecules and
the activity level of molecules. Thus we need to have a thorough understanding of the chemical
behavior of acids and bases in solution. In the final module of this unit you will see these
concepts in action in protein ligand binding.
pH
The hydrogen ion concentration, [H+] of a solution is an
important property because biological systems contains functional groups
whose properties are changed by changes in the hydrogen ion
concentration.
Since the hydrogen ion concentations are usually much less than
one, and can vary over many orders of magnitude, a different scale is
used for the hydrogen ion concentration, the pH scale. The pH is the
-log of the proton concentration:
The log conversion reduces a 10 fold change in hydrogen ion
concentration to a one unit change in pH. The minus sign changes the
negative numbers that would be obtained from log[H
+] to
positive ones. Since the pH scale is an inverse scale the
concentration of protons is high at low pH and low at high pH. A
solution is said to be acidic if the pH is less than seven, and basic if
the pH is above 7. A solution is neutral if its pH is equal to 7.0.
The image below shows the pH of a number of common fluids.
pH of various compounds.
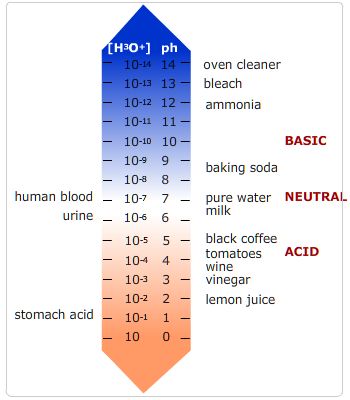 On the left are biological compounds and on the right are some foods and cleaning products.
On the left are biological compounds and on the right are some foods and cleaning products.
Differences in hydrogen concentration
Acid Dissociation and pH
For our studies, the Bronsted definition of an acid will be used, we will define an acid as a proton donor and a base
as a proton acceptor. Hydrochloric acid, like sodium chloride, is a
strong electrolyte because it completely dissociates in aqueous solution
into charged ions. Hydrochloric acid is also a strong acid because
when it completely dissociates it also completely donates all of its
protons.
Many molecules are weak electrolytes and exist in an equilibrium (indicated by in the general equation below) between the starting molecule and its dissociated parts. Thus dissociation can be seen as an acid (HA) in equilibrium with a proton (H+) and the corresponding conjugate base (A-).
In general:
Specifically for Acetic acid:
Acid Dissociation and Equilibrium
In this, and following, sections, we will begin to develop a
quantitative relationship between the strength of an acid and how the
acids ionization state will depend on the pH of the solution. When
placed in water, acetic acid dissociates into the conjugate base,
acetate, and a proton.
The extent to which the acid will dissociate in pure water is expressed as K
a, the equilibrium constant for dissociation of an acid:
Ka is the ratio of the mathematical product of
the concentration of each product of the reaction (in this case the
charged species) to the concentration of the reactants (in this case the
neutral species). The square brackets [ ] around the terms indicate
concentration and is usually expressed in molar concentration
(moles/Liter). In the Learn By Doing you will explore the equilibrium of
an acid dissociation in pure water. In this simulation you can assume
the concentration is molecules per beaker and thus the concentration
will be equal to the total number of molecules in the experiment.
Ka is constant for an acid.
In the tutorial you learned that the equilibrium constant Ka is the
same for an acid regardless of the starting concentration. Ka
describes the relationship between the concentration of the protonated acid and
the charged dissociated species at equilibrium. You also saw that even though
Ka is constant, the values of the concentration of each species
was varying over time. After an initial rapid dissociation of the acid, an
equilibrium was reached. At equilibrium, the number of individual molecules will
fluctuate, but on average, a balance, or equilibrium mixture, is maintained. The
balance of products and reactants is defined by Ka.
Each acid has its own Ka
The equilibrium constant, Ka, is a property of the molecule and its
environment. The chemical nature of the molecule has the largest effect on
Ka. Secondary to this, the environment of the group can shift the
Ka. Therefore, every acidic group has its own Ka value.
For the purposes of this course, we will use the approximation that Ka
is the same for all molecules with the same functional group. The table below lists the
Ka values for several acids.
| Ka for various acids |
|---|
| Acid |
Ka
|
| Acetic Acid |
0.0001 |
| Boric Acid |
0.000000001 |
| Ammonia (NH4+) |
0.000000001 |
|
Relationship of Ka and pKa
The Ka is a small number, usually much less than 1, and can vary by
several orders of magnitude from compound to compound. For example, acetic acid
has a Ka of around 0.0001 or 1.0 x 10-4 while the
Ka of ammonia is 105 fold smaller. Due to the wide
range of Ka, it is more practical to represent the Ka on a
minus log scale, in the same way as the hydrogen ion concentration.
Consequently, the negative log of the Ka is used to represent acidity
constants:
pKa = -log(Ka)
A change in the pKa from 5 to a pKa 4 is the change by a
factor of 10 in the Ka. The table below will help you see the
relationship between Ka and pKa. The pKa is a
convenient scale for comparing the dissociation constants of weak acids because
the pKa scale is similar to the pH scale. The relationship between the two will
be explained in the next few pages. Remember that the pKa is
a property of the acidic group while pH is a property of the solution.
| Ka
|
pKa
|
| 0.1 |
1 |
| 0.01 |
2 |
| 0.001 |
3 |
| 0.0001 |
4 |
| 0.00001 |
5 |
Just as the pH scale indicates the relative proton concentration of various
solutions, the pKa indicates the relative strengths of the different
acids. If a reaction has a large equilibrium constant, then the concentration of
the products will be higher than the reactants. In the case of the acidity
constant, a larger Ka indicates a more completely dissociated acid,
or a stronger acid. Since the pKa = - log (Ka), strong
acids will have small pKa values.
For this course you do not need to memorize the pKa values for all of
the different groups. The values will always be given to you, or you can find
them in the Functional Groups Glossary. However, it is useful to remember the
following pKa values:
| Acidic Group |
pKa |
Example |
| -COOH (carboxyl) |
4.0 |
Aspartic acid side chain. |
| -NH3 (amino) |
9.0 |
Lysine side chain |
Control of Acid Dissociation
Placing acetic acid in pure water establishes an equilibrium between the charged
and neutral forms according to the unique dissociation constant of the acid.
This occurs because acetic acid is the only proton donor in the solution. What
would happen if protons are added to the solution from another source, for
example a strong acid, or a biochemical reaction that releases protons? Answer
the question before continuing.
Because the equilibrium constant Ka does not change, the presence of
additional protons from another source must shift the ratio of acetate ions to
acetic acid, and a new equilibrium point would be reached.
For the following acid dissociation:
with an equilibrium
constant of:
How will the equilibrium position change when the proton
concentration changes? If protons are added [A
-] must decrease and
[HA] increase in order for K
a to remain the same. Likewise if protons
are removed by adding a strong base then [A
-] must increase, and [HA]
decrease to keep K
a constant. The ratio of [A
-] to [HA]
changes depending on the total [H
+] of the solution.
To quantify how the equilibrium point is affected by pH, it is instructive to
rearrange the equilibrium equation to generate the Hendersson-Hasselbach
equation. First we start with the general equilibrium equation for
Ka:
and then take the -log of both sides of the equation.
You may recognize that -log(K
a) is pKa and the
-log([H
+]) is pH. By substituting these into the equation we have
the Hendersson-Hasselbach equation.
This equation can be used to calculate the ratio of
[A
-]/[HA] for any weak acid, provided the pK
a of the acid
and the pH of the solution are known.
Effect of pH on Protonation State
The fraction of a group that is protonated: [HA]/{[HA]+[A-]}
can be calculated at any pH using the Hendersson-Hasselbach equation.
Such a plot is shown below for acetic acid. Note that:
- When pH = pKa, one half of the molecules will be
protonated
- When the pH is one unit lower than the pKa 90% of the
molecules will be protonated.
- When the pH is one unit higher than the pKa only 10% of
the molecules are protonated.
- For pH values outside +/-1 pH unit from the pKa the group
is essentially fully protonated (pH << pKa) or
fully deprotonated (pH >> pKa)
 The fraction protonated as a function of pH is plotted for a
weak acid with a pKa=4.0, such as acetic acid. Note that
when the ph=pKaone-half of the molecules will be
protonated. More than half are protonated with the pH is lower than
the pKa while less than half are protonated when the pH
is greater than the pKa.
The fraction protonated as a function of pH is plotted for a
weak acid with a pKa=4.0, such as acetic acid. Note that
when the ph=pKaone-half of the molecules will be
protonated. More than half are protonated with the pH is lower than
the pKa while less than half are protonated when the pH
is greater than the pKa.
Buffers
Weak acids can act as pH buffers when the pH is within approximately one
unit of the pKa for the acid. A buffer solution will resist
changes in pH as a strong acid or base is added to the solution. The
following shows the pH of a solution of a weak acid as a strong base,
e.g. hydroxide, is added or titrated into the solution.
 The pH of a solution of a weak acid with a pKa of
4.0 is shown as base is added to the solution. The initial pH of the
solution is low, and most of the weak acid is fully protonated. As
base is added the weak acid remains protonated, consequently the
added base causes a rapid rise in pH due to the neutralization of
protons. When the pH of the solution is near the pKa of
the weak acid, the acid begins to dissociate, producing protons that
neutralize the added base. Since the base is being neutralized the
pH climbs more slowly. When the pH is above the pKa most
to the weak acid is deprotonated and can no longer provide protons
to neutralize the base, consequently the pH climbs rapidly again.
The pH region that is within one unit of the pKa is
considered to be the buffer region.
The pH of a solution of a weak acid with a pKa of
4.0 is shown as base is added to the solution. The initial pH of the
solution is low, and most of the weak acid is fully protonated. As
base is added the weak acid remains protonated, consequently the
added base causes a rapid rise in pH due to the neutralization of
protons. When the pH of the solution is near the pKa of
the weak acid, the acid begins to dissociate, producing protons that
neutralize the added base. Since the base is being neutralized the
pH climbs more slowly. When the pH is above the pKa most
to the weak acid is deprotonated and can no longer provide protons
to neutralize the base, consequently the pH climbs rapidly again.
The pH region that is within one unit of the pKa is
considered to be the buffer region.
Understanding the reverse titration, the addition of a strong acid to a
solution of conjugate base follows the same logic. At high pH, the added
protons do not protonate the weak acid since the pH is much higher than
the pKa. In the buffer region of pH = pKa+/-1, the
protons that are added to the solution will not decrease the pH, instead
they will convert some of the conjugate base, A-, to the acid
form, HA. At the lower edge of the buffer region the weak acid is almost
fully protonated (pH=pKa-1), consequently it cannot absorb
any additional protons.
The buffering of pH plays an important role in the normal function of the
cell. If the pH drops too low, or becomes too high, the cell can no
longer function. The principal buffer that is used in biological systems
is the carbonic acid, a weak acid with an effective pKa of
6.4, ideally suited to act as a buffer when the pH ~ 7.0
So far, we have discussed the major elements and functional groups
that are important in the
functioning of a cell. Together these elements and functional groups
define the major properties of the four classes of macromolecules that
make up a cell: carbohydrates, proteins, lipids and nucleic acids. In
this module, we will explore two of these classes: carbohydrates and
proteins.
Carbohydrates, proteins and nucleic acids are all examples of
polymers. These polymers are very large molecules composed
of smaller units joined by covalent bonds using a common set of
chemical reactions. Proteins are linear polymers of amino acids
all joined by peptide bonds. Polysaccharides are the carbohydrates
joined through
glycosidic bonds in sometimes quite complex branched structures.
Later in the course you will encounter DNA and RNA which are polymers of
nucleic
acids linked by phosphodiester bonds. This unit includes a discussion
of the structures of polymers of carbohydrates and of amino acids.
Carbohydrates
One of the simplest of the biological molecules is the carbohydrate. The name is
descriptive of the character of this class of molecules since they all have the general
formula of a hydrated carbon.
We are starting with this class of molecules because they are the basis for the classical naming of the enantiomers encountered in
biological molecules. But before we examine the structure of these compounds, let us
look at some of the basic uses/functions of the carbohydrates.
The primary function of carbohydrates is as a source of energy.
You will recall that molecules
are a collection of atoms connected by covalent bonds. In general,
single covalent bonds
can be represented as having approximately 100 Kcal/mole of energy
associated with the
force that holds the two atoms together. The most common carbohydrate
in nature is glucose, which has the general formula (C(H2O))6 or C6H12O6.
When glucose is completely metabolized in a cell, 673 kcal of energy
is released for each mole of glucose. The net equation for this process
of glucose oxidation can be written as follows:
The challenge for the cell is to capture released energy and convert
it to a useful form so that it can do work (e.g. power movement or
build new macromolecules). How cells manage to do this is discussed in
the Metabolism section of the course.
The second function for the carbohydrates is structure. In this case, structure is not only
what a polymer of the carbohydrates has, but it is also what that structure contributes
to the cell. For example, cellulose is a linear polymer of glucose that interacts with
other cellulose polymers to form fibers that interact to form the basic structure of the
cell wall of plants.
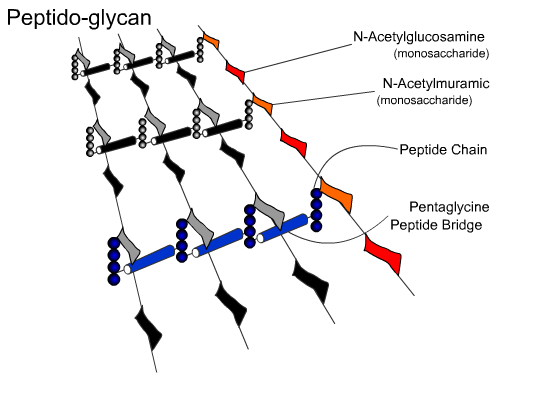
Another example is the peptido-glycan that makes up the cell wall structure of a
bacterium. Peptido refers to a peptide, which is a fragment of a protein or a short
polymer of amino acids, and glycan refers to a polysaccharide, a polymer of
carbohydrates. In this case, the polymer of carbohydrates includes building blocks other
than glucose, but the end result is the formation of a matrix. In both cases the
resulting fibers and matrices provide scaffolding that gives rigidity (structure) and
protection to the organisms with which they are associated.
A third function for carbohydrates is that of cell recognition and signaling. Just
as we identified a peptido-glycan as a conjunction (conjugation) of a peptide with a
polysaccharide, other complex carbohydrates are conjugated to other molecules to form
glycoproteins (carbohydrates linked to proteins) and glycolipids (carbohydrates linked
to lipids). Because a very large number of structures can be made from a few
monosaccharides (simple carbohydrates), a very large number of different structures can
be made from a few simple carbohydrates, as will be seen later. This large number of
different structures can therefore be used as unique signals for identification of
individual cell types.
Monosaccharides
The simplest of the carbohydrates fall into two categories or structures that differ only
in the arrangement of the atoms as seen below. In fact, in biological systems it is
quite easy to convert between the two forms using catalysts. The unique functional
group associated with that category defines each of the categories.
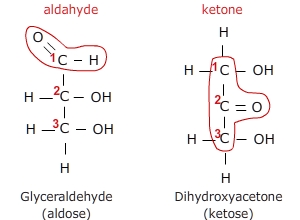
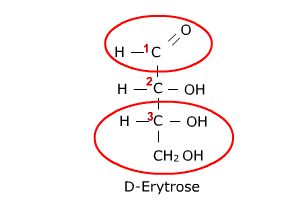
In the case of the structure on the left, the number 1 carbon (the top carbon) contains
the carbonyl that is flanked by a hydrogen and a carbon thus making this an aldehyde.
This category of carbohydrates is, thus, referred to as aldoses. In contrast, the
structure on the right has its carbonyl at the number 2 carbon (the center carbon) and
this carbonyl is flanked by carbons on both sides, thus, making this carbonyl a ketone.
Carbohydrates containing this ketone group are referred to as ketoses.
- Aldoses
-
(Definition)
A category of simple carbohydrates where the number 1 carbon (the top carbon)
contains the carbonyl that is flanked by a hydrogen and a carbon thus making
this an aldehyde.
Glyceraldehyde was chosen as the first carbohydrate to study because it provides another
example of the bioselectivity of biological systems. In this case the selectivity is
based on structural discrimination and will apply to all of the carbohydrates.
Examine the glyceraldehyde structure given above and determine
which of the carbons has a unique composition of four bonding partners.
The number 1 carbon has three bonding partners: an oxygen that is double
bonded to the carbon, a hydrogen, and the remainder of the structure.
The number 2 carbon has an aldehyde, a hydrogen, a hydroxyl group and
the rest of the molecule attached, while the number three carbon has a
hydroxyl, two hydrogens and the upper portion of the molecule attached.
The number 2 carbon isthe only carbon that has its four bonds involved
with four different groups and, thus, is a unique carbon identified as a
chiral or asymmetric center. Based on the fact that this chiral carbon
has tetrahedral bonding structure, it can form structural enantiomers.
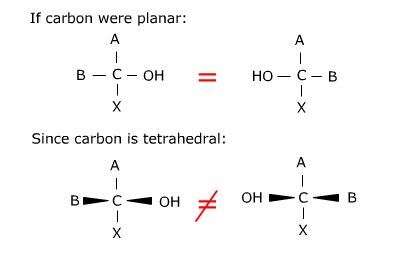
You should convince yourself that it is not possible to superimpose the bottom two
structures. They are in fact different structures. For glyceraldehyde, where A is an
aldehyde, B is a hydrogen and X is the hydroxymethyl (CH2OH) group, the structure on the
left would be referred to as D-glyceraldehyde and the structure on the right would be
referred to as L-glyceraldehyde based on the orientation of the hydroxyl functional
group.
Biological systems are selective and while it is possible to
synthesize both of the
glyceraldehyde enantiomers in the chemistry lab, a cell primarily
produces and uses the
D form of glyceraldehyde. This is dictated by the way in which
molecules are selected
and used in a cell. Every reaction in a cell is catalyzed by an
enzyme and these enzyme
catalysts have the ability to discriminate between different
structures. In this case
the enzymes can discriminate between D and L carbohydrates. Thus as a
general rule, all carbohydrates in biological systems are D.
Just as the cell can recognize the difference between D and L, the
expectation for this
course is that you are able to recognize the structures of the
carbohydrate but it will not be necessary to know how to draw the
structures.
Aldoses
Now we will build larger aldoses by adding one carbon at a time to the structure. In
doing this, two components will remain unchanged, the aldehyde group will always be the
number one carbon and the bottom or last two carbons will always represent the D form of
the carbohydrate (monosaccharide).
Add another carbon to the structure given above. How many possible structures result
from adding another carbon to this carbohydrate? What must remain constant in the
representation of these structures?
One of these structures is very important to the biological system: D-ribose. This
aldose makes up the backbone structure of RNA and a derivative of it, 2-deoxyribose,
makes up the backbone of DNA.
Similarly, an additional carbon can be added to the pentoses to
form the possible
hexoses (six carbon monosaccharides). An expansive tree on structures
can be built
starting with the simplest aldose. While all of these structures are
possible starting
with D-glyceraldehyde, the biological system only utilizes a few of
these structures. Of
the hexoses, the predominant compound is D-glucose, one of the most
prevalent sugars in biology. The linear structure of glucose is shown
below, along with ribose.
It is useful to provide shorthand notations and alternative ways of
representing the carbohydrates. Below are equivalent alternative representations of the
glucose and ribose. The form on the right is typically found in books and manuscripts. It
should always be remembered that while the structure on the right would appear to
represent carbon as having planar bonding, in fact each of the carbons has a tetrahedral
bonding structure.
Cyclization of Aldoses
The five (ribose) and six (glucose) membered aldoses have been
depicted in their linear form. However, these compounds will
spontaneously form 5 member or 6 member ring structures if
possible. These are more stable forms of the compounds.
Now let’s represent the ribose structure as it is actually found in solution. As
previously mentioned, compounds will spontaneously form 5 or 6 member rings if possible.
For aldoses this is possible because of the reactive character of the aldehyde group.
For ribose this means that if the oxygen on the number 4 carbon forms a bond with the
number one aldehyde carbon and the hydrogen on the hydroxyl shifts to the carbonyl
oxygen on the aldehyde the result is a five-member ring referred to as a furanose.
Examination of the result of this transformation, in which the total number of carbons,
hydrogens and oxygens between the two structures has not changed, shows that the number
one carbon has now become a new asymmetric (chiral) center where it wasn’t before the ring
closure. This new chiral center is called the anomeric carbon. You should examine this structure and
convince yourself that the four different substituents attached to the anomeric carbon are
unique, making it a chiral center.
The drawing below shows not only the possible structures resulting from the ring closure
but also shows the more traditional representation of these structures in their ring
configuration. In all of these cases the structure is still ribose. The two new
structures created by the ring closure create two additional conformations of the number
one carbon: alpha with the hydroxyl on the right of the stick structure or pointing down
in the ring representation, and beta with the hydroxyl on the left of the stick
structure and pointing up in the ring structure. It should also be noted that as
represented in the drawing these structures are all in equilibrium with each other and
the alpha structure can be converted to the beta structure and visa versa as long as
each structure can be converted to the free aldehyde structure.
The hexoses are capable of forming a six member pyranose
ring by the same mechanism. Ring formation in glucose is shown in the
diagram below. As with ribose, a new chiral center is formed, giving
two possible forms of glucose, alpha and beta.
Epimers: Monosaccharides that differ by chirality at one
position (besides the anomeric carbon) are called epimers. For example,
glucose and galactose are epimers of each other. Galactose is one of
the monosaccharides that make up lactose, or milk sugar. There are
enzyme catalysts capable of inter-converting galactose to glucose by
inverting the chiral center. Thus the galactose that is released from
lactose can be concerted to glucose and used for energy.
Ketoses
- Ketoses
-
(Definition)
A ketose is a carbohydrate with a carbonyl at the number 2 carbon that is
flanked by carbons on both sides. This is a polar,
hydrophilic, water-soluble molecule.
In the same way that the larger aldoses were generated, single carbons can be added to
the structure of the dihydroxyacetone and an expansive tree of structures
result. If we examine the result of adding a single carbon, the result is the acquisition
of an new asymmetric center. In the case of a four carbon ketose, which is formed by the addition of a C(H2O)
group below the ketone group in dihydroxyacetone phosphate, the new
chiral center can be either D or L. In general, only the D structures
of the ketoses are used by biological
systems due to bioselectivity.
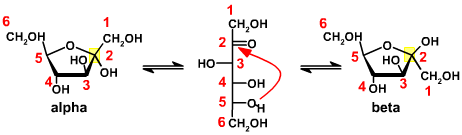
While it is clear that many ketose structures are possible, only a few are used in
biological systems and the focus for this course will be on the hexose D-fructose. The
fructose structure has the ketone group as at the number 2 carbon and the D
designation based on the chirality at carbon number 5. Applying the principle that compounds
will spontaneously form stable ring structures, and using a mechanism
similar to that employed for the aldoses, fructose will spontaneously form a five-member ring
structure. These structures are depicted below and are in equilibrium with
each other through the free ketone structure.
While the monosaccharides can serve directly as sources of energy, as will be seen
elsewhere in the course, they are also the building blocks for many molecules that are
used for structure, energy storage and signaling. Here we will explore the process by
which simple carbohydrates are linked together using a condensation reaction. Time will
be spent on understanding this reaction since it is the same reaction that is used to
link amino acids together to make proteins and fatty acids to glycerol to made the components of biological membranes.
As described previously the aldehyde form of aldoses and ketone form of ketoses
spontaneously form five and six member ring structures called furanoses and pyranoses
respectively. This ring closure was an example of an alcohol functional group reacting
with the carbon of the carbonyl functional group in either the aldehyde (aldose) or
ketone (ketose) group. This is illustrated below. Notice that while atoms are moving
from place to place on the structure, there is no net gain or loss of atoms in the
closing of the ring to form what is referred to as a hemiacetal. Furthermore,
this ring closure is freely reversible, which allows the alpha form of the ring to be in
equilibrium with the beta form.
Starting with the hemiacetal (closed ring) structure, the unique, newly created,
asymmetric center has a special name: the anomeric carbon. In the structure of glucose
given below, the anomeric carbon, which is created by forming the six member
(pyranose) hemiacetal structure, is highlighted. You should examine this structure and
convince yourself that the four different substituents attached to that carbon are
unique .

This anomeric carbon is the target for the formation of a covalent bond between it and
potentially any hydroxyl functional group on any other monosacchardie. The covalent bond
formed between the anomeric carbon and a hydroxyl group is called a glycosidic bond
and the final structure is referred to as an acetal. The equation for the
formation of the glycosidic bond is given below. In this reaction, a molecule of water
is lost during the combining of the two glucose molecules to form the glycosidic bond.
This type of reaction is referred to as a condensation
reaction. The reverse of this reaction, requiring the addition of a water
molecule, is referred to as a hydrolysis reaction. These combined forward and reverse reactions (condensation and
hydrolysis) form the basis for the creation of most of the covalent assemblies in
biological systems.
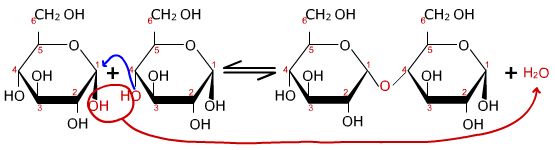 The hydroxyl groups from the 1 carbon and 4 carbon react to produce an α 1,4-glycosidic bond and water.
The hydroxyl groups from the 1 carbon and 4 carbon react to produce an α 1,4-glycosidic bond and water.
While all linkages between the anomeric carbon of one carbohydrate
and every hydroxyl of a second carbohydrate are possible, it is
important to understand, that in biological systems, very little is left
to chance, and in fact, every reaction that takes place in a cell is
catalyzed by an enzyme. Just as we
have stated that bioselectivity dictates that D carbohydrates are
used by biological
systems, the formation of glycosidic bonds is catalyzed by specific
enzymes that direct
the formation of specific glycosidic bonds between the anomeric
carbon of one defined
carbohydrate and a specific hydroxyl function of a defined second
carbohydrate.
Each
resulting disaccharide is a different structure. This is another
example of
bioselectivity.
It should also be noted that while the transformation from the alpha to the beta form of
an individual carbohydrate at the anomeric carbon is freely reversible and dictated by
equilibrium, that freedom of conversion is lost once the anomeric carbon is involved in
a glycosidic bond. Furthermore, the condensation/hydrolysis reaction is generally
written as an equilibrium; however, the glycosidic (acetal) linkage is very stable and
does not spontaneously break (hydrolyze) without the input of energy and the use of a
specific enzyme. The stability of the glycosidic bond contributes to the effective use
of the polysaccharides in maintaining structure.
Carbohydrates generally exist as di- and polysaccharides used in transport, energy
storage, structure and signaling. While the glycosidic bond is always between the
anomeric carbon of one carbohydrate and the hydroxyl of another carbohydrate, the number
of hydroxyls per monosaccharide and the different orientations of the hydroxyl on the
anomeric carbon make the number of potential structures extremely large. Here we will
focus on some of the biologically important di- and polysaccharides with an emphasis on
the ability to identify and describe the structures not the ability to draw the
structures.
Disaccharides
Two of the most common disaccharides are lactose, or milk sugar, and
sucrose, or common table sugar. Each is a form of carbohydrate
storage and represents an example of a hetero-disaccharide, i.e. disaccharide formed from
two different monosaccharides.
In the case of sucrose, the composition is a molecule of glucose and a molecule of
fructose. The structure is described as glucose-alpha,beta
1,2-fructose. In this structure it is instructive to identify the anomeric carbon on the
glucose and the fructose molecules since they are both involved in the formation of the
glycosidic bond.
A third disaccharide that provides a transition to the common
polysaccharides is maltose, a homo-disaccharide. The structure is a
homo-dimer made up of two molecules of glucose linked by an alpha-1,4 glycosidic bond.
Maltose is commonly found as a dietary supplement and naturally is produced as an
intermediate breakdown product of starch. As such it also represents the basic repeating
unit of the polysaccharides starch and glycogen.
Polysaccharides
Homopolysaccharides: Three major homopolymers found in nature are starch, glycogen and cellulose. All three are made of the same building block (subunit or monomer) – glucose, but they are different in structure and function.
Starch: The structure of starch has two components: amylose and
amylopectin. Amylose is a linear polymer of glucose subunits linked
end-to-end by alpha-1, 4 glycosidic bonds.
Amylopectin has the same backbone polymer structure as amylose, but also
contains branches from the backbone linked by alpha -1,6 glycosidic
bonds every 20-30 glucose subunits along the amylose backbone. This
gives rise to a highly branched structure used in the storage of glucose
in plants.
Glycogen is another example of a homo-polysaccharide of glucose
with a repeating structural unit of glucose. For glycogen, there is no
linear form, but there is a highly branched form that resembles
amylopectin with branch points separated by only 8-10 glucose subunits
along the backbone. This creates a much denser, more highly packed
structure used for storage of glucose in mammalian cells.
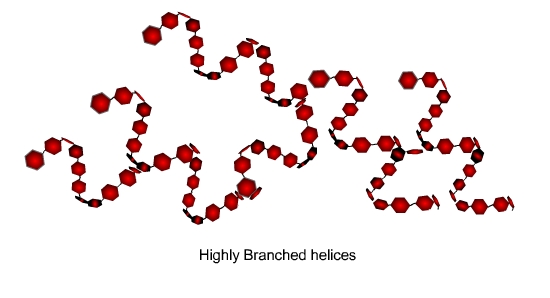
Cellulose plays a structural role as the key component of a plant
cell wall. Cellulose is an unbranched polymer, analogous to amylose;
however, the linkages between the glucose subunits are beta-1,4, rather
than alpha-1,4. This difference in configuration of glycosidic bond
leads to differences in structure and in function. Beta-1,4 linkage
results in the linear and extended shape of the polymer structure, which
allows the strands of cellulose to align and form a hydrogen-bonding
network between the hydroxyls on the individual glucose subunits. This
hydrogen bonding adds to the rigidity of cellulose cell wall.
Another demonstration of bioselectivity is at play in the
discrimination between cellulose and amylose by our digestive system.
The structural difference between the beta-1,4 link in cellulose and the
alpha-1,4 glycosidic bond in amylose is sufficient to require different
enzymes to break (hydrolyze) the bonds separating the glucose subunits
in each structure. The difference is significant because it prevents
humans from being able to use cellulose as a glucose source, while we
are able to use glycogen and starch. Humans do not posses the enzyme
that hydrolyzes the beta-1,4 link in cellulose.
Hetero-Polysaccharides: An enormous set of possibilities for structures exist
if one considers that variations can occur between the orientation of the
glycosidic bond (alpha vs. beta), the sites to which the bond can be made on an
adjacent monosaccharide, the number of different
carbohydrates involved (glucose, galactose, ribose, etc), and the order in which they
appear.
Such variation, even when a few monosaccharides are included, can give rise to an enormous
number of uniquely recognizable structures. In some cases these varied structures can be
attached to proteins and lipids to be used as identification/signaling devices on the
surface of cells. Each cell type and each species of microorganism can display a unique
identification on its surface to be used in cell recognition and to identify partners in
cell-cell interactions.
Many other hetero-polysaccharides exist to carryout a variety of functions within an
organism. A unique hetero-polysaccharide produced by bacteria is the peptido-glycan that
forms the basic structure of the bacterial cell wall. In this case there is a linear
hetero-polysaccharide that acts much like the linear strands of cellulose in the plant
cell wall. However, as implied by the name, the peptido-glycan is a covalent complex
between a polysaccharide (glycan) and a peptide (a fragment of a protein). In the
bacterial peptido-glycan, the linear hetero-polysaccharide chains are linked together by
covalent bonds using a peptide as the linker. As seen in the illustration below, this
means that the bacterial cell wall is fully connected by covalent bonds while the plant
cell wall is stabilized by hydrogen bonding between hydroxyl groups on parallel strands.
Introduction
Amino acids are the building blocks of proteins. The sequence of amino acids in
individual proteins is encoded in the DNA of the cell. The physical and chemical
properties of the 20 different, naturally occurring amino acids dictate the
shape of the protein and its interactions with its environment. Certain short
sequences of amino acids in the protein also dictate where the protein resides
in the cell. Proteins are composed of hundreds to thousands of amino acids. As
you can imagine, protein folding is a complicated process and there are many
potential shapes due to the large number of combinations of amino acids. By
understanding the properties of the amino acids you will get an appreciation for
the limits of protein folding and how to predict the potential higher order
structure of the protein.
|
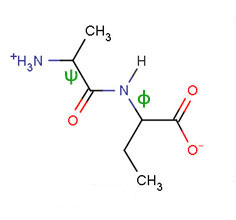
The numbering of the carbon atoms in an amino acid follows the nomenclature used for carboxylic acids.
The first carbon adjacent to the carboxyl group is the α-carbon,
followed by the β-carbon, etc. as illustrated in the diagram to the
right. All amino acids that are found in proteins have the same backbone
structure, the acidic carboxylic acid group, the α–carbon, and an
amino group that is attached to the α-carbon,
hence the name α-amino acids. The sidechain, or R, group is attached
to the α-carbon and is different for each amino acid.
Note that the α proton is often not drawn, but its existence should
be inferred from the fact that carbon forms four single bonds.
The amino group has a pKa value of ~9, thus it is protonated at pH 7.0. The carboxylic acid group has a pKa of 2.0,
and thus it is deprotonated at pH 7.0. Each of the 20 amino acids has a different sidechain(R group),
which is attached to the α–carbon. Use the exercise below to identify these important functional groups in an amino acid.
|
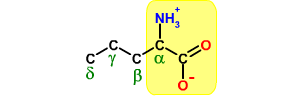 General structure of an α-amino acid. The mainchain atoms, which are found in all amino acids, are highlighted.
The remaining carbon atoms (β, γ, etc) form the sidechain of the amino acid. These differ from one amino acid to the next. General structure of an α-amino acid. The mainchain atoms, which are found in all amino acids, are highlighted.
The remaining carbon atoms (β, γ, etc) form the sidechain of the amino acid. These differ from one amino acid to the next.
|
Chirality: Because there are four different groups attached to the central carbon, the
α–carbon is an asymmetric or chiral center. The chiral center gives rise to D and
L enantiomers for each amino acid. However, where bioselectivity dictated the
dominance of D carbohydrates, the same discrimination process for amino acids
gives rise to the dominance of the use of L amino acids in nature.
If all of the amino acids have the same basic structure with an amino, a
carboxyl and a hydrogen fixed to the α–carbon, then the large variation in
the properties and structure of the amino acids must come from the fourth group
attached to the α–carbon. This group is referred to as the side chain
of the amino acid or the R group. The structures of
the 20 common amino acids are shown below. Clicking on the amino acid
in the table, displays three-dimensional model of that amino acid below
the table.
The sidechain groups of these amino acids contain many of the same functional
groups that were discussed in the first module and can be found in the
Functional Groups. To learn more about the amino acids, watch the video of Dr.
Bill Brown's lecture on amino acids.
It may be helpful to review the Functional Groups by clicking the Learn-By-Doing
link below.
Peptide Bond
Proteins are polymers of amino acids. The amino acids are joined together by a
condensation reaction similar to that described for the formation of the
glycosidic bond in polysaccharides. Each amino acid in the polymer is referred
to as a residue. Individual amino acids are joined together by the attack of the
nitrogen of an amino group of one amino acid on the carbonyl carbon of the
carboxyl group of another to create a covalent peptide bond and yield a molecule
of water as shown below.
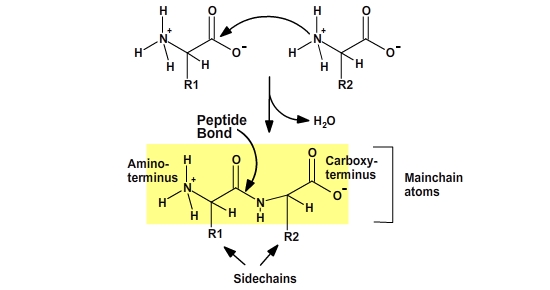 Peptide bond formation occurs by a dehydration reaction. The amino
group of the second amino acid attacks the carbonyl carbon of the first,
forming the peptide bond and releasing water. The resultant dipeptide has an
amino terminus (left) and a carboxy-terminus (right). The mainchain atoms,
which are the same for each residue in the peptide, include the nitrogen and
its proton, the α–carbon and its hydrogen, and the C=O group. The R-groups form the
sidechain atoms.
Peptide bond formation occurs by a dehydration reaction. The amino
group of the second amino acid attacks the carbonyl carbon of the first,
forming the peptide bond and releasing water. The resultant dipeptide has an
amino terminus (left) and a carboxy-terminus (right). The mainchain atoms,
which are the same for each residue in the peptide, include the nitrogen and
its proton, the α–carbon and its hydrogen, and the C=O group. The R-groups form the
sidechain atoms.
The resulting peptide chain is linear, defined by the mechanism that builds the
polymer, and has defined ends. Short polymers (< 50 residues or amino
acids) are usually refereed to as peptides, and longer polymers as proteins.
Because the synthesis takes place from the alpha amino group of one amino acid
to the carboxyl group of another amino acid, the result is that there will
always be a free amino group on one end of the growing polymer (the N-terminus)
and a free carboxyl group on the other end (the C-terminus). Note that the
potential exists for the formation of amide (peptide) links involving the
carboxyl and amino groups in the side chains, but bioselectivity directs the
synthesis to be linear, involving only the alpha amino and alpha carboxyl
groups.
Note that after the amino acid has been incorporated into the protein, the
charges on the α–amino and the carboxyl groups have disappeared. Thus the mainchain
atoms have become polar functional groups. Since each residue in a protein has
exactly the same mainchain atoms, the functional properties of a protein must
arise from the different sidechain groups.
By convention, the sequences of peptides and proteins are written with the
N-terminus on the left and the C-terminus on the right. The name of the
N-terminal residue is always the first amino acid. The name of each amino acid
then follows. The primary sequence of a protein refers to its amino
acid sequence.
Protein sequence
A protein is composed of amino acids attached in a linear order. This basic level of protein
structure is called it's primary structure and derives
from the formation of peptide
bonds between the individual amino acids.
The order of the amino acids is determined by information encoded
in the cell's genes. An
example of a protein sequence is shown below where the one letter
abbreviations are used for each of the 20 amino acids used in cellular
protein synthesis.
Amino acid sequence of
Human Estrogen Receptor
 Amino acids are
indicated using the single letter code. For example, the amino acid glycine is abbreviated with the letter G.
Amino acids are
indicated using the single letter code. For example, the amino acid glycine is abbreviated with the letter G.
Higher order structure is determined by the Primary Structure
Proteins do not exist as linear threads in the cells but rather as
spontaneously folded higher order structures. The higher order
structure is determined by the amino acids in the
primary structure. The function of a protein is determined by its higher
order structure.
Proteins take on structure in stages defined by the interactions
of amino acids with each other in the primary sequence. The higher order
stages of secondary, tertiary, and quaternary structure of the
estrogen receptor are shown in the movie below. Although the exact
structure of the protein was determined
using biophysical methods employing sophisticated instruments and
calculations, the ability to predict the three dimensional structure
(tertiary structure) of a protein de novo is not yet possible. It is
possible to make excellent predictions of secondary structure from
primary structure and the use of measured constraints on structure based
on the structure of the peptide bond and characteristics of individual
amino acids is making it possible to approach predictive methods.
The stages or levels of protein structure are displayed here:
-
Primary Structure: The amino acid sequence of the protein, with no regard for the conformation of the amino acids.
-
Secondary Structure: short range interactions involving only mainchain atoms resulting in α-helices and β-sheets
-
Tertiary Structure: long range interactions resulting in the 3-D Folding of a single polypeptide chain.
-
Quaternary Structure: The interaction of two or more peptide
chains to make a functional protein. The oxygen transport protein
hemoglobin shows this level of structure. The functional protein is
composed of four chains.
Constraints Determining Secondary Structure
Peptide Bond is Planar
When two amino acids are joined a peptide bond is formed
through a
condensation reaction. The carbonyl electrons are partially shared by
the
carbon nitrogen peptide bond giving the carbonyl carbon/amide nitrogen
bond a slight double bond character. Because of this partial double bond
character, rotation around the C-N peptide
bond is prevented and thus the peptide bond is planar. This is the first
and a major constraint placed on protein folding. The figure below
depicts the six atoms that are included in the planar peptide bond: the
carbonyl carbon, the carbonyl oxygen, the alpha carbon attached to the
carbonyl carbon, the amide nitrogen (of the second amino acid), the
hydrogen attached to the amide nitrogen, and the alpha carbon of the
second amino acid. Two possible orientations of the planer peptide
bond are possible, the trans from and the cis form. Although both are
planer, the trans form is more stable and this form is shown in the
diagram below.
Since there is no rotation around the C-N bond of the peptide
bond, the only possible freedom of rotation in an amino acid residue is
the carbonyl carbon-alpha carbon single bond, which is denoted as
psi(Ψ), and the amide nitrogen-alpha carbon single bond, which is called
phi(Φ). If one knows all of the phi-psi angles of rotation for every
amino acid residue in a protein, it is possible to define the secondary
structure of that protein. It should be noted that not all phi-psi
angles are possible due to steric interference, electronic repulsion by
side chains of the same charge and other factors all of which represent
constraints in the folding of a protein.
Explaining Φ and Ψ
Watch this short video for a description of the Φ and Ψ angles.
Side Chain Constrains Φ and Ψ Angles
Secondary structure is generally defined by the interaction
of amino acids adjacent to each other in the primary sequence. Elements
of secondary structure include the α-helix, the beta-sheet and the
beta-turn. These structures involve short range interactions that
involve hydrogen bonding between peptide bonds and do not involve the interactions of amino acid side chains.
Alpha Helix One of the constrained pairs of Φ-Ψ angles
gives rise to the secondary structural element referred to as the
α-helix. The α-helix is the prominent
structure seen hemoglobin shown previously. Because of the regular
nature of the structure, every Φ-Ψ angle pair in an α-helix is the same.
α-helical structure
is stabilized by hydrogen bonding between the backbone carbonyl oxygen
and the amide hydrogens. In α-helices there are 3.7 amino acids per turn
and the carbonyl of the n-th amino
acid hydrogen bonds to the amide hydrogen of the n+4 amino acid. The
hydrogen bonds are parallel to the long axis of the helix and the side
chains extend to the outside of the helical cylinder.
Beta-Sheet The second predominant secondary structure
element is the
β-sheet. When the Ψ and Φ angles are close to 180 degrees the
peptide chain is fully extended, the β-strand conformation exists.
A β-sheet forms when several β-strands run
either parallel or anti-parallel to each other. The structures are
stabilized by hydrogen bonds between the strands and the anti-parallel
structure appears to be more stable due to maximization of the number of
possible hydrogen bonds. The atoms participating in hydrogen bonding in
a β-sheet are the same as those for α-helices: the carbonyl oxygen and
the amide hydrogens. In the case of a β-sheet the hydrogen bonds are in
the plane of the sheet and perpendicular to the peptide chain. The side
chains project above and below the plane of the β-sheet.
Proline Kinks the System
Proline is unusual among the amino acids and is technically
referred to as an imino acid. It is a cyclic molecule forming a five
member ring that closes the linear hydrocarbon side chain with the
α-amino group. Because of this there is no rotation about the Ψ bond and
therefore, proline terminates helix formation and places further
constraints on the folding of the peptide chain.
Lecture on Protein Structure
View the following video describing protein structures.
Hydrophobic interactions drive tertiary structure formation
Proteins contain a mix of
hydrophobic and
hydrophilic amino acids.
Hydrophobic molecules contain no or very few highly electronegative atoms and do not
form strong bonds with water. In aqueous solutions hydrophobic molecules are driven
together to the exclusion of water. This is an example of the hydrophobic effect previously
described in the section on bonding.
As a protein folds to its final three-dimensional structure, the hydrophobic parts of the
protein are forced together and away from the aqueous environment of the cell.
Stabilization of the Folded State: The tertiary structure is
determined by non-covalent interactions that involve amino acid side chains
(side chain and backbone interactions, or side chain-side chain interactions).
Amino acid residues involved in these interactions can come from distant parts
of the polypeptide chain bringing the chain into a more compact shape. The
non-covalent interactions between the side chains include the following:
- Iconic bonds
- Hydrogen bonds
- Hydrophobic/van der Walls interactions
Your should explore the Jmol structure to determine which of the
following is more important in stabilizing the tertiary structure.
Disulfide bonds stabilize tertiary structure
Protein folding and stabilization is driven primarily by non-covalent
interactions. α-helices and β-sheets are stabilized by hydrogen
bonding. Tertiary structure is driven and stabilized by hydrophobic interactions
and involves all types of non-covalent bonding. As you will see in the next
page, quaternary structure is stabilized primarily by all types of
complimentary, non-covalent interactions. One covalent bond is possible and is
formed when two cysteine amino acids (sulfhydryl containing side chain) are
close together as a result of tertiary structure formation. The sulfhydryl group
is highly reactive and will covalently bond with another sulfhydryl group to
form a covalent disulfide bond (a cystine residue). Disulfide bonds also form
between peptide chains in forming quaternary structure, for example in the
structure of antibodies. The disulfide bond is reversible and is sensitive to
the environment.
More than one peptide forms a functional protein
There are many examples of proteins that require more than one peptide chain to be
functional. In the case of the estrogen receptor two identical peptide chains come
together to form the functional protein. This is called a homo dimer. Hemoglobin is
composed of four peptide chains; two identical alpha chains and two identical beta chains.
The individual peptides or sub-units of estrogen and hemoglobin are held together by all
of the non-covalent bonding types:
hydrogen bonding,
ionic bonding and
hydrophobic interactions
.
The interaction of the subunits in a quaternary complex
represents another form of equilibrium. The individual polypeptide
chains are made separately and must associate through specific
complimentary interactions.
Antibodies represent a different type of quaternary structure
because the individual subunits are linked by disulfide bonds. The
structure is a dimer of dimers and the basic unit is a large peptide
(the heavy chain) bound to a smaller chain (light chain) by disulfide
bonds. Since this dimer is a combination of two different peptides it is
called a heterodimer. Two of these heterodimers are joined by disulfide
bonds between the heavy
chains to form the final tetrameric structure.
Protein - Ligand Interations
In this module we will use all of the concepts previously introduced in the section on
biological chemistry to study the interaction, or binding, of small molecules to
proteins. If the small molecule binds to a protein, but undergoes no further changes,
then it is referred to as a ligand. If the molecule binds to the protein, and
then is changed by a protein-catalyzed chemical reaction, the small molecule is referred
to as a substrate, the resultant altered compound is the product, and
the protein becomes an enzyme. Although ligands and substrates are usually
small molecules, proteins themselves can be ligands and substrates for other
proteins/enzymes.
Protein-ligand interactions serve several important functions in the cell. They may be
responsible for buffering the ligand concentration and the transport of ligands. Oxygen
binding to myoglobin is an example of buffering while the protein hemoglobin is involved
in the transport of oxygen from the lungs to the tissues.
It is also possible that binding the ligand causes the functionality of the protein to
change from active to inactive, or from inactive to active. For example, the estrogen
receptor protein, shown below in yellow and purple, binds DNA only when estrogen (red)
is bound to the protein. Estrogen is the "key" that turns the estrogen receptor "on." In
the figure below, the estrogen receptor is depicted with its secondary structure
highlighted. As you can see it is mostly α-helices. The estrogen receptor
represents a quaternary structure composed of a dimer of two protein chains shown as
yellow and purple in the figure. 17-β-estradiol (an estrogen), the red
space-filling balls, fits in a properly sized pocket on each of the proteins in the
dimer. Only when both pockets of the dimer are occupied by estrogen is the receptor
activated and able to bind to DNA. As you can see the estrogen binding pocket is only a
small part of the total protein size.
The estrogen receptor binds to estrogen and this causes estrogen receptor to bind to DNA
and turn genes on and off. The result in some cells is the commencement of cell division. When estrogen
falls off, estrogen can no longer bind DNA and division may stop. If estrogen did not
dissociate from the estrogen receptor then cell division could not be controlled and
cancer could result.
Energetics of Protein-Ligand Complex Formation
The binding of ligands and substrates relies primarily on the characteristics of the
non-covalent bonds and the functional groups associated with both the protein and the
small molecule. The interactions that can be used are:
- van der Waals interactions: the protein and its ligand are
complementary in shape, optimizing the contact between the two surfaces.
- hydrogen bonding
- hydrophobic interactions,
- ionic bonds
The functional groups that are responsible for these interactions
are those found in the
side chains of amino acids, the peptide bonds in the protein and the
small molecule
binding partner. Following some simple rules it is possible to
predict how two
molecules might interact. In some cases It is also possible to
predict how a cell might alter
physiological conditions to regulate that binding. You will explore
that interactions that stabilize bound estrogen to the estrogen
receptor in the following activity:
Ligand Binding is Reversible
Because protein and ligand interactions are non-covalent in nature it is possible for the
ligand to dissociate from the protein. This reversible binding is critical and is
generally represented as:
Macromolecule (protein) + Ligand <=> [Macromolecule Ligand]
M + L <=> [ML]
The bound complex is surrounded by brackets, [ ], to indicate that the ligand is
physically associated with the macromolecule.
In the following pages you will explore reversible binding and how we can determine the
strength of the interaction between a protein and its ligand by experimental
measurements.
As discussed previously, proteins bind and release their ligands. In this section we will
explore the reversible binding and the equilibrium state. We will use myoglobin, as an
example, to explore the equilibrium established between the bound protein ligand complex
and free reactants. Myoglobin is a single polypeptide chain protein that binds oxygen to
create an oxygen buffer in muscle cells. The reversibly binding of oxygen to myoglobin
can be expressed as follows:
By reversibly binding oxygen, myoglobin acts as an oxygen buffer. When free oxygen is high
myoglobin binds oxygen and the equilibrium shifts to the right. If the oxygen
concentration falls the equilibrium shifts to the left to free oxygen and protein. The
change in equilibrium is only possible because protein ligand binding is reversible. The
direction of the equilibrium, either towards more complex or towards free reactants,
depends primarily on the amount of free oxygen in the cell. Oxygen supply is always
changing depending on activity level. Myoglobin supply is essentially constant in the
cell since protein synthesis is relatively a much slower process.
In the Learn By Doing below, you will explore the equilibrium between free protein,
ligand, and the complex, and how ligand concentration affects the equilibrium.
The number of the contacts between a protein and its ligand and the environment of the
binding pocket determine how tightly the ligand binds to the protein. It is difficult to
predict how tightly a ligand will bind to a protein or to predict the structure of
ligands that will bind tightly a binding pocket of a known protein. There is intensive
research in this area and forms the basis for many computational studies used in drug
discovery laboratories. The tightness of the binding is a measurable parameter and
describes the affinity of a ligand for a protein.
In the previous section the binding of oxygen to myoglobin was defined by the
equation:
This equation describes an association of
two molecules. In the previous analysis of weak electrolytes we wrote the dissociation
of a weak acid as the taking apart of a molecule that was bound by a weak covalent bond.
While there are no covalent bonds being formed during the association of oxygen and
myoglobin, there is the same type of equilibrium established between the association of
myoglobin and the dissociation of the weak acid. Thus, where we used a Kd (K) as a
constant that defined the strength of the weak electrolyte's ability to retain its
proton, it is also possible to define the affinity of the ligand for the protein by a
constant referred to as the equilibrium constant, Keq. For the association reaction as
it is written above, the equilibrium constant would be expressed as follows:
In general the affinity of the ligand will increase proportionately as the number of
specific interactions between the ligand and the protein increase. In the example of the
binding pocket of the estrogen receptor there are several hydrogen bond contacts and a
hydrophobic pocket for the mostly hydrophobic estrogen. Other protein ligand binding
interactions are based on fewer contacts, as in the case of oxygen binding to myoglobin
and thus the affinity is lower.
Previously, you explored how proteins interact with their ligands, using the example of
myoglobin and oxygen. Proteins are not only storage proteins, but can also act as
hormone receptors as seen with the estrogen receptor and many serve as enzymes that
catalyze processes in the cell.
- Catalyst
-
(Definition)
A catalyst is a participant in a chemical reaction that speeds up the reaction
but is not consumed itself. Enzymes are biological catalysts that mediate the
conversion of substrate to product, by lowering the activation energy of the
reaction. Enzymes re not consumed or modified during the process. Enzymes are
generally made of proteins but nucleic acid enzymes, ribozymes, have also been
discovered.
General Reaction
Cells operate within the laws of chemistry and physics, and utilize only
chemical reactions that occur naturally.
Many of
these reactions are extremely slow, requiring years to eons to complete.
For example, your favorite book contains paper made of cellulose
from trees. This paper will take decades to degrade in the environment and
mammalian systems lack the enzyme that is specific to degrade cellulose,
however, a bacteria found in the stomach of cows and other ruminants have the
ability to digest cellulose and provide glucose to the cow. For a cow to survive
it cannot wait decades to digest a meal of hay. To meet the speed of life,
chemical reactions in cells are sped up by enzyme catalysts.
Enzymes bind to substrates in a manner similar to the way myoglobin binds oxygen
or the estradiol binds to the estrogen receptor, but enzymes can go one step
further. In this case the ligand is specifically referred to as the
substrate (the molecule that the enzyme will convert to product)
and it binds to a specific binding region of the enzyme referred to as the
active site. Once bound, the ligand, or substrate, can either
simply reversibly come off the enzyme, or it can be converted into a new
compound or product. The general form of the reaction is:
Where E = enzyme, S = substrate, ES = the enzyme•substrate complex, and P =
product. The square brackets around the ES complex indicate that the complex is
inferred, because it is very unstable and generally not directly measurable.
Catalysts Lower Activation Energy
Enzymes and other catalysts lower the activation energy for converting a
specific substrate to product, as shown in the graph below. The activation
energy is the energy required for a chemical reaction to proceed to product. By
lowering the activation energy, a chemical reaction can proceed much more
quickly. Note that enzymes have no effect on the free energy of the reaction.
Enzymes are not consumed in the process, but are recycled for further
catalysis.
Catalysts Lower Activation Energy
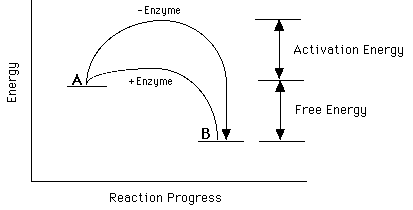 This graph shows how enzymes catalyze reactions by lowering the
activation energy. Without a catalyst, the amount of energy required for the
reaction is large. By fixing a substrate in its active site, enzymes lower
the activation energy, increasing the rate of the reaction.
This graph shows how enzymes catalyze reactions by lowering the
activation energy. Without a catalyst, the amount of energy required for the
reaction is large. By fixing a substrate in its active site, enzymes lower
the activation energy, increasing the rate of the reaction.
The Rate of Product Formation.
In the last section, the action of an enzyme was described as taking place in two
parts. The first part involves the binding of substrate to enzyme in a
reversible equilibrium as modeled in the way of ligand binding. The second part
involves the conversion of the [ES] complex to product and the release of the
product from the enzyme. This step represents the catalysis step in the enzyme
catalyzed reaction. These parts are illustrated in the overall reaction below:
The rate, or velocity (v), of the reaction is usually determined by measuring the
amount of product (P) produced/unit time:
The dependence of the reaction velocity on the kinetics of substrate binding,
substrate release, and conversion to product is difficult to calculate under all
conditions. However, if steady-state conditions are assumed then a very
simple relationship can be found. Steady-state conditions imply that during the
measurement the concentrations of the various species, e.g. [ES] ,do not change.
Under steady-state conditions the following equation describes the relationship
between the measured rate (v) of the reaction and the initial concentration of substrate
(S) and the concentration of the enzyme. This equation is the Michaelis
Menton-Briggs Haldane equation.
In this equation, kcat is the rate at which the [ES] complex decays
to product. Vmax is kcat multiplied by the total
amount of enzyme, Et. It is therefore the maximum velocity at which
the reaction can occur under a fixed enzyme concentration. Km is also
a constant that reflects the affinity of the enzyme for the substrate and is an
approximation of the dissociation equilibrium constant. Both kcat and
Km depend on the particular enzyme and substrate.
This equation can also be represented graphically by the hyperbolic curve
illustrated below. This curve can be dissected into two parts: a region at the
beginning where the velocity of the reaction increases proportionately with the
amount of substrate present initially in the reaction, and the later region of
the curve, at high substrate concentration, where the rate of the reaction is
independent of the substrate concentration. This latter region defines
saturation of the enzyme by substrate and the velocity approaches the value of
Vmax.
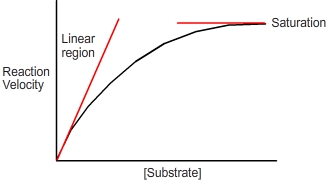 Effect of substrate concentration on enzyme velocity.
Effect of substrate concentration on enzyme velocity.
Enzyme catalysis is affected by many factors including substrate and
enzyme concentrations, pH, temperature, and the presence of inhibitors.
The learn-by-doing on this page will explore the effect of substrate
concentration. The following page will explore the effect of enzyme
concentration, pH, temperature, and inhibitors on the velocity of the
enzyme catalyzed reaction.
Enzyme Catalysis
This simulation will demonstrate the effect of substrate on the reaction
velocity.
Left alone, enzymes would deplete a cell of all the substrates. In some cases, this may
be desirable, such as enzymes that degrade old, worn-out proteins. In other cases, there
may be multiple uses for a substrate and depletion, by one enzyme, may be detrimental to
the working of a cell. In some cases, such as the digestion of food, the enzyme needs to
be functional in the gut but not in the cell. Thus, several types of regulations have
evolved that the cell can use to control enzyme rate.
Control by Enzyme Concentration
A common way of regulating enzyme activity is by increasing or decreasing
the amount of enzyme by regulation of the rates of synthesis in the cell.
If you recall, the rate of reaction is proportional to the concentration
of the enzyme-substrate complex ([ES]). The concentration of this complex
can be increased in two ways: i) either by increasing [S], which you
explored on the previous page, or ii) by increasing the total amount of
enzyme in the cell.
Control by Temperature
One mechanism to control the rate of product formation is through temperature.
Velocity of product formation increases linearly as the temperature increases
until a critical temperature is reached as indicated in the following graph.
Velocity of Product formation versus temperature.
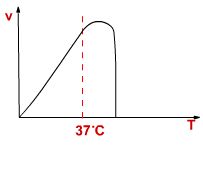
Product formation ceases beyond the critical point due to two effects of
temperature. One effect is to decrease the non-covalent interactions between
enzyme and substrate. Remember that ionic and hydrogen bonds are weak
non-covalent bonds representing 5 Kcal/mole each.only 5 kcal/mol in strength.
When the temperature increase matches the energy of the bond, the energy cn be
used to break the bond. Thus if the enzyme cannot bind the substrate, no product
is formed.
The second effect of temperature is to cause proteins to denature (unfold).
Denaturation means that the protein loses the secondary, tertiary, and
quaternary structure required for function. As discussed earlier, the protein
tertiary structure is stabilized by non-covalent bonding. As with substrate
binding, increased temperature can provide the energy necessary to break the
weak non-covalent bonds. In some cases, the beaking of a single hydrogen bond
can alter the structure of the enzyme sufficiently to cause the loss of function
(i.e. loss of the ability to bind substrate, to recognize the substrate or to
catalyze product formation).
Control by pH
The pH of the environment a protein finds itself in can drastically affect the
proteins function and denaturation state. Have you ever put lemon and cream into
hot tea at the same time? The cream curdles because the low pH of the lemon
juice causes the proteins to denature. pH also affects the ability of a protein
to bind to its ligand. At low pH hydrogen bond acceptors such as carboxyl groups
become protonated. If they are protonated then they cannot form the critical
ionic bonds between the protein and the ligand. The graph below shows the
activity of a typical protein as a function of pH. This protein is most active
around neutral pH.
The cell can use pH as a mechanism to control enzyme action. Enzymes used in
digestion that destroy other proteins could be detrimental to a cell. However,
typically these enzymes are only active at low pH such as is found in the
stomach or in intracellular compartments such as the lysosome that is maintained
at lower pH than the cytoplasm.
In vivo Control - Competitive and Non-competitive Inhibition
Temperature and pH represent environmental means of regulating the velocity of an
enzyme catalyzed reaction. However, in a typical organism (in vivo) the
temperature and pH are usually regulated and therefore cannot be used to control
enzyme activity. Consequently, the velocity of enzymatic reactions are usually
regulated by molecular interaction of a second molecule with the enzyme. These
are generally termed inhibitors if the binding of the second molecules causes a
loss of enzyme activity (reduced velocity at any given substrate concentration).
In competitive inhibition the second molecule binds in the active site
thus preventing the substrate from binding. The regulatory molecule competes
with the substrate for the active site. This decreases the rate at which the
enzyme can convert the substrate to product because the enzyme spends part of
the time unable to bind the substrate. This can be seen in the graph below
plotted as the dashed red line. The occupancy of the binding pocket is in
equilibrium between the binding of the substrate or the inhibitor. As you can
see from the curve the maximum velocity is unchanged, but the substrate
concentration required to reach maximum velocity is increased.
Non-competitive inhibition involves a second small molecule binding to the
enzyme. In this case there are two binding sites: one for the substrate and a
distinct one for the inhibitor. When the inhibitor binds to the enzyme, the
substrate binding pocket changes such that the substrate is no longer able to
bind. It may also be possible for both the substrate and inhibitor to bind the
enzyme at the same time but the enzyme is unable to complete the conversion of
substrate to product. Non-competitive inhibition results in a lower maximum
velocity for the enzyme when the inhibitor is present as can be seen in the
green curve. Inihibitors of this type are often referred to as
allosteric inhibitors because they cause the shape (steric) of the
enzyme to assume a different (allo) form.
Introduction to Lipids
The cell is composed of two distinctive environments: the hydrophilic aqueous
cytoplasm and the hydrophobic lipid membranes. The lipid environment is defined
by the family of molecules that are characterized by their hydrophobic nature
and their common metabolic origin. Three members of the lipid family of
molecules will be discussed in this course: fats (triacylglcerol),
phospholipids, and steroids.
The Structure of Lipids
Lipid molecules are slightly soluble to insoluble in water.
Lipids are hydrophobic because the molecules consist of long, 16-18
carbon, hydrocarbon chains (or backbones) with only a small amount of
oxygen containing groups. Lipids serve many functions in organisms. Fats
(or triglycerides) are used to store energy. Phospholipids and steroids
are the key components of cell membranes. Lipds are also the major
components of waxes, pigments, and steroid hormones.
Fats (triacylglycerols, triglycerides)
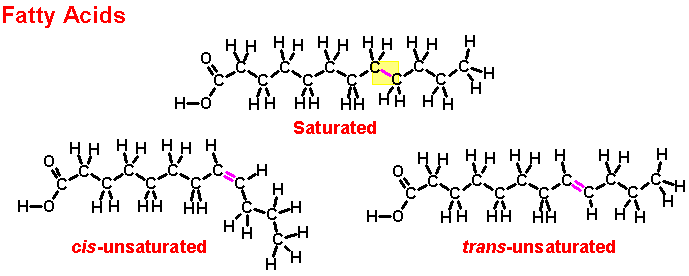 Various
forms of a 12-carbon fatty acid are shown. The top structure is the
fully saturated fat, lauric acid. The two lower structures show two
forms of unsaturated fatty acids that can be generated from lauric acid
by formation of a single double bond. The cis (left) form is typically
found in biological lipids, note that the double bond has kinked the
hydrocarbon chain. The trans (right) form is generated by a chemical
process that transforms unsaturated fatty acids to saturated fatty
acids. This process is used to convert liquid vegetable oils to solid
margarine.
Various
forms of a 12-carbon fatty acid are shown. The top structure is the
fully saturated fat, lauric acid. The two lower structures show two
forms of unsaturated fatty acids that can be generated from lauric acid
by formation of a single double bond. The cis (left) form is typically
found in biological lipids, note that the double bond has kinked the
hydrocarbon chain. The trans (right) form is generated by a chemical
process that transforms unsaturated fatty acids to saturated fatty
acids. This process is used to convert liquid vegetable oils to solid
margarine.
Fats are synthesized from two different classes of molecules:
fatty acids and alcohol. The fatty acids are unbranched hydrocarbons
that terminate with a single carboxyl functional group. The fatty acids
are generally 16-22 carbons long and can be both saturated and
unsaturated. Saturated fatty acids have no carbon-carbon double bonds
(they are saturated with hydrogen), while the unsaturated fatty acids
have one to three double bonds along the backbone carbon chain. These
double bonds introduce "kinks" in the carbon chain which have important
consequences on the fluid nature of lipid membranes. Unsaturated fatty
acids have lower melting points than saturated fatty acids.
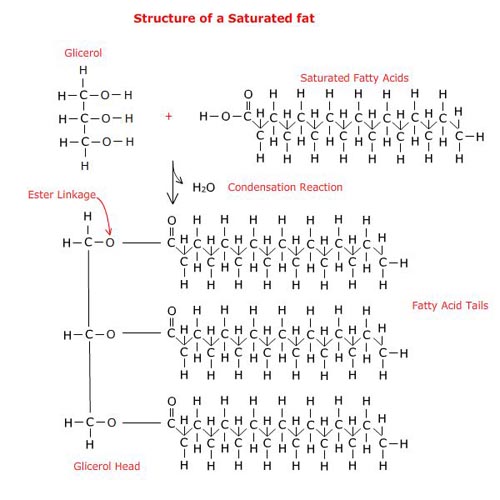
To construct a fat, or triacylglycerol, three fatty acid molecules are
attached to the glycerol through an ester bond between the carboxyl
group of the fatty acids and the three alcohol groups of a glycerol
molecule. This is another example of a condensation reaction that
results in formation of an ester in this case and the release of a water
molecule. A fat molecule can be composed of one, two, or three different
types of fatty acids each of which can be saturated or unsaturated.
An unsaturated fat has at least one unsaturated fatty acid, whereas a
saturated fat has none. Because the double bonds of the unsaturated
fatty acids introduce kinks in the hydrocarbon backbone, unsaturated
fats will not pack into a regular structure and thus remain fluid at
lower temperatures. A saturated fat will pack well and be a solid a low
temperatures.
Fats are mainly energy storage and insulating molecules. Per gram, fats
contain twice as much energy as carbohydrates. Layers of fat also
surround the vital organs of animals to cushion them, and layers of fat
under the skin of animals provide insulation.
Phospholipids
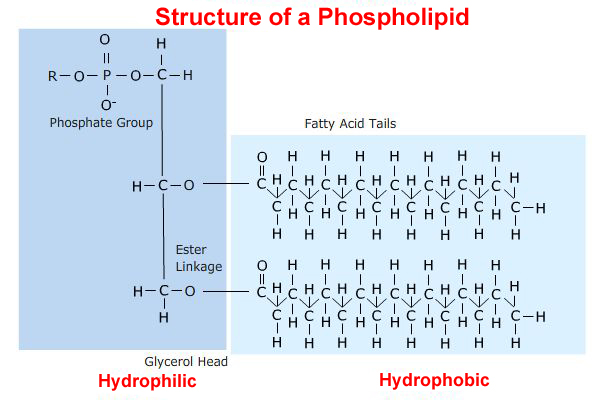
Phospholipids contain only two fatty acids attached to a glycerol head.
This occurs by a condensation reaction similar to the one discussed
above. The third alcohol of the glycerol forms an ester bond through
reaction with phosphoric acid. This is another example of a condensation
reaction between an acid and an alcohol with the release of water. As a
triprotic acid (i.e it has three acidic functions on the phorphorus
atom) the phosphate group attached to the glycerol has the potential to
form ester links with a variety of other molecules such as
carbohydrates, choline, inositol and amino acids. The phosphate group
along with the glycerol group make the head of the phospholipid
hydrophilic, whereas the fatty acid tail is hydrophobic. Thus
phospholipids are amphipathic:a molecule with a polar end and a
hydrophobic end. When phospholipids are in an aqueous solution they will
self assemble into micelles or bilayers, structures that exclude water
molecules from the hydrophobic tails while keeping the hydrophilic head
in contact with the aqueous solution. View the animation that
demonstrates the formation of micelles and bilayers.
Click the green arrow to play the animation.
Phospholipids serve a major function in the cells of all organisms: they
form the phospholipid membranes that surround the cell and intracellular
organelles such as the mitochondria. The cell membrane is a fluid,
semi-permeable bilayer that separates the cell's contents from the
environment, see animation below. The membrane is fluid at physiological
temperatures and allows cells to change shape due to physical
constraints or changing cellular volumes. The phospholipid membrane
allows free diffusion of some small molecules such as oxygen, carbon
dioxide, and small hydrocarbons, but not charged ions, polar molecules
or other larger molecules such as glucose. This semi-permeable nature of
the membrane allows the cell to maintain the composition of the
cytoplasm independent of the external environment.
A closer view of a Lipid Bilayer forming a membrane
Steroids
The steroids are a family of lipids based on a molecule with four fused
carbon rings. This family includes many hormones and cholesterol.
Cholesterol is a component of the cell membrane in animals and functions
to moderate membrane fluidity because it restricts the motion of the
fatty acid tails.
Structure of Cholesterol
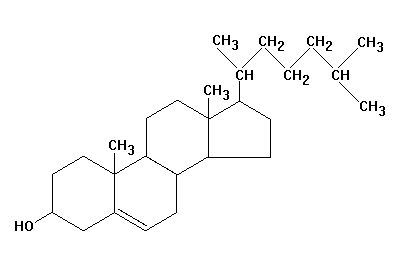
Cholesterol in the membrane decreases the fluidity.
Review of Lipids
Examine the effect of cholesterol on membrane fluidity using the animations below.
Use the play/pause button on the left to start or stop the
animation. Use menu to move between scenes.
The Structures of the Cell Membrane
Fluid Quality of Membranes
The cell membrane must be a dynamic structure if the cell is to grow and
respond to environmental changes. To keep the membrane fluid at
physiological temperatures the cell alters the composition of the
phospholipids. The right ratio of saturated to unsaturated fatty acids
keeps the membrane fluid at any temperature conducive to life. For
example winter wheat responds to decreasing temperatures by increasing
the amount of unsaturated fatty acids in cell membranes. In animal cells
cholesterol helps to prevent the packing of fatty acid tails and thus
lowers the requirement of unsaturated fatty acids. This helps maintain
the fluid nature of the cell membrane without it becoming too liquid at
body temperature. The fluidity of the membrane is demonstrated in the
following animation. The lipids in the membrane are in random bulk flow
moving about 22 µm (micrometers) per second. Phospholipids freely move
in the same layer of the membrane and rarely flip to the other layer.
Flipping of phospholipids from one layer to the other rarely occurs
because flipping requires the hydrophilic head to pass through the
hydrophobic region of the bilayer.
Click the green arrow to play the animation.
The Mosaic Quality of Membranes
Proteins
Because the cell membrane is only semipermeable, the cell needs a
way to communicate with other cells and exchange nutrients with
the extracellular space. These roles are primarily filled by
proteins. Membrane proteins are classified into two major
categories, integral proteins and peripheral proteins. Integral
membrane proteins are those proteins that are embedded in the
lipid bilayer and are generally characterized by their
solubility in non-polar, hydrophobic solvents. Transmembrane
proteins are examples of integral proteins with hydrophobic
regions that completely span the hydrophobic interior of the
membrane. The parts of the protein exposed to the interior and
exterior of the cell are hydrophilic. Integral proteins can
serve as pores that selectively allow ions or nutrients into the
cell. They also transmit signals into and out of the cell.
Unlike integral proteins that span the membrane, peripheral
proteins reside on only one side of the membrane and are often
attached to integral proteins. Some peripheral proteins serve as
anchor points for the cytoskeleton or extracellular fibers.
Proteins are much larger than lipids and move more slowly. Some
move in seemingly directed manner while others drift.
Carbohydrates
The extracellular surface of the cell membrane is decorated with
carbohydrate groups attached to lipids and proteins.
Carbohydrates are added to lipids and proteins by a process
called glycosylation, and are called glycolipids or
glycoproteins. These short carbohydrates, or oligosaccharides,
are usually chains of 15 or fewer sugar molecules.
Oligosaccharides give a cell identity (i.e., distinguishing self
from non-self) and are the distinguishing factor in human blood
types and transplant rejection.
Membranes are Asymmetric
As discussed above and seen in the picture, the cell membrane is
asymmetric. The extracellular face of the membrane is in contact
with the extracellular matrix. The extracellular side of the
membrane contains oligosaccharides that distinguish the cell as
self. It also contains the end of integral proteins that
interact with signals from other cells and sense the
extracellular environment. The inner membrane is in contact the
contents of the cell. This side of the membrane anchors to the
cytoskeleton and contains the end of integral proteins that
relay signals received on the external side.
Summary: Membranes as Mosaics of Structure and Function
The biological membrane is a collage of many different proteins embedded in the
fluid matrix of the lipid bilayer. The lipid bilayer is the main fabric of the
membrane, and its structure creates a semi-permeable membrane. The hydrophobic
core impedes the diffusion of hydrophilic structures, such as ions and polar
molecules but allows hydrophobic molecules, which can dissolve in the membrane,
to cross it with ease. Proteins determine most of the membrane's specific
functions. The plasma membrane and the membranes of the various organelles each
have unique collections of proteins. For example, to date more than 50 kinds of
proteins have been found in the plasma membrane of red blood cells.
Membrane Transport
The cell membrane provides a semi-permeable barrier between the inside and the outside of
the cell. This barrier provides control for transport of nutrients, ions and signals
between the highly variable outside environment and the relatively well-defined interior
of the cell. This unit of the course will explore the ways in which molecules can pass
across the membrane (diffusion and active transport), can be transported into the cell
without passing across the membrane (endocytosis), and can send signals for actions
within the cell without actually passing across the membrane themselves (signal
transduction).
Passive/Simple Diffusion:
Both large and small molecules follow the same general principal of
diffusion. Molecules
spontaneously move from areas of high concentration to areas of low
concentration
following Brownian motion. The classic example is the diffusion of a
drop of ink placed
in a beaker of water. The concentrated drop of color slowly disperses
(diffuses) until
at some point equilibrium is reached in which the beaker appears to
have a uniform
color. The following animation depicts this simple diffusion process.
Add ink to each beaker and watch the diffusion process. After a period
of time, is there a difference in the distribution of ink in each
beaker? Follow the yellow ink molecule in each beaker for some time. In
which simulation does the yellow ball take longer to transverse the
beaker?
Question: If this simulation represented the movement of the
ink in air (no gravity considerations), would the movement be faster or
slower in water? Why?
The introduction of a cell or liposome into the solution places a barrier to the molecules.
As three different molecules diffuse to equilibrium they encounter the lipid bilayer
depicted by the horizontal membrane across the center of the stage in the following
animation. Note that one type of molecule passes freely through the lipid bilayer while
the second type of molecule only occasionally passes through the membrane and the lipid
bilayer is totally impermeable to the third type of molecule.
Of the first two types of molecules, the first type might include a molecule such as
cholesterol which has some solubility in water but is highly soluble in the non-polar
environment of the lipid bilayer and thus will freely pass into the hydrophobic
environment of the membrane, distribute freely in the membrane and then some fraction
will dissolve in the aqueous environment of the cytoplasm. A second example of this type
of permeable molecule is water which while polar is small and able to freely pass across
the membrane. The lipid bilayer is much less permeable to the second type of molecule
indicating a more polar character and a larger size. Examples of such molecules are the
sodium and chloride ions. As a general rule, charged molecules are much less permeable
to the lipid bilayer. The third type of molecule is very polar, in many cases
charged, and usually a larger molecule. Examples are carbohydrates and amino acids.
Aspirin
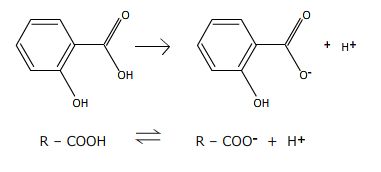 In Protonated and non-Protonated States
In Protonated and non-Protonated States
Osmosis
Cells continually encounter changes in their external ionic environment and will
spontaneously respond by attempting to equalize the concentration of ions on the inside
and outside of the cell. Because the plasma membrane (lipid bilayer) is significantly
less permeable to ions than water, the establishment of an equal concentration of the
ions on either side of the membrane is accomplished by the net movement of water toward
the higher concentration of ions to reduce the concentration. This movement of water in
response to an imbalance of solute (ion) is referred to as osmosis. This is illustrated
in the following simulation.
Three different conditions may exist in the relationship between the solute (ion)
concentration and solvent (water) concentration across a membrane. Isotonic, hypertonic
and hypotonic refer to the relative concentration of the solute (small molecules) in the
extracellular (outside) space surrounding the cell relative to the solute concentration
inside the cell. In an isotonic solution, the concentration of the solute and therefore
solvent water (water potential) is the same on both sides. A hypotonic solution is one
whose solute concentration is lower (water concentration is higher [i.e. high water
potential]) in the extracellular space than inside the cell. Because the water (the
solvent) can more easily pass through the membrane than can the solute (ions), the net
flow is spontaneous in the direction of the solvent (water) moving from its higher
concentration (high water potential) outside the cell to the inside of the cell.
Conversely, a hypertonic solution refers to an extracellular solution with a higher
solute concentration (lower water concentration [i.e. low water potential]) outside the
cell than inside the cell. In this case, the more permeable solvent, water, would flow
spontaneously out of the cell toward the low water potential to dilute the solute
molecules and create an equal concentration of solute molecules on both sides of the
membrane.
Facilitated Diffusion
Cells must be able to move polar molecules such as nutrients and ions across the lipid
bilayer of the membrane in order to carry out life processes. But the molecules will
still move spontaneously down a concentration from high to low concentration. To allow
the polar molecules, which are not soluble in the lipid bilayer, to pass across the
hydrophobic barrier it is necessary to provide ports, channels or holes through the
membrane. These channels can either allow the molecules to move freely according to
their concentration differences or they can be gated channels that control the movement
of the polar molecules according to the needs of the cell. In most cases these channels
are very discriminatory and will only allow specific molecules to pass; another example
of bioselectivity. The channels facilitate the movement of molecules that otherwise
would not be spontaneously permeable to the lipid bilayer. The process of moving
impermeable molecules across a membrane using channels or pores is referred to as
facilitated diffusion. Because the molecules are moving down a
concentration gradient the process is driven by simple diffusion. The following
simulation depicts the facilitated diffusion of glucose across the membrane using the
glucose permease transporter.
In some cases it is necessary to move molecules against a gradient.
The eukaryotc cell, a
typical mammalian cell, has many compartments within the cell each
surrounded by a lipid
bilayer membrane. In most cases the environment within the
compartment is different than
that in the cytoplasm. An example is the lysosome, a degradative
organelle (membrane
bound compartment within the cell) whose function is to digest
macromolecules delivered either from the outside of the cell or from
other compartments within the cell. To
carry out this function the lysosome maintains a much lower pH inside
the lysosome
relative to the cytoplasm. At equilibrium, the concentration of
protons would be equal on
both the inside and outside of the lysosome.
To decrease the pH inside of the lysosome, the concentration of protons will need to be
greater inside the lysosome than in the cytoplasm. To accomplish this protons will need
to move from a low concentration to a high concentration. This is a non-spontaneous
process and requires the cell to do work to move the ions up-hill against the gradient.
To do work, the cell must expend energy and actively move (pump) the ions. This process
is referred to as active transport. The source of energy for this process in most
biological systems is the hydrolysis of ATP.
The following animation depicts another example of active transport;
the sodium-potassium ATPase. This active transport system moves three
sodium ions out of the cell and two potassium ions into the cell, each
against a gradient.
Transport Proteins
Facilitated diffusion and active transport both require channels or ports in the membrane
through which the generally non-permeable molecules can pass. These protein transporters
contribute to the mosaic character of the fluid mosaic character of the biological
membrane. There are a variety of different structures associated with transport proteins
and at the same time many transport proteins that carry out similar functions (e.g. ion
channels) have structural similarities while maintaining their ability to discriminate
between molecules. Thus transport proteins have been classified both by structure and by
function. For the purposes of this course, the classification will be that of function
though similarities in structure will be observed in the examples chosen.
There are three classifications of transport proteins based on mechanism of transport: Uniport,
Symport and Antiport. The animations on the following pages will demonstrate the three classes of
proteins with examples of each.
Transport Proteins
Facilitated diffusion and active transport both require channels or ports in the membrane
through which the generally non-permeable molecules can pass. These protein transporters
contribute to the mosaic character of the fluid mosaic character of the biological
membrane. There are a variety of different structures associated with transport proteins
and at the same time many transport proteins that carry out similar functions (e.g. ion
channels) have structural similarities while maintaining their ability to discriminate
between molecules. Thus transport proteins have been classified both by structure and by
function. For the purposes of this course, the classification will be that of function
though similarities in structure will be observed in the examples chosen. There are
three classifications of transport proteins based on mechanism of transport: Uniport,
Symport and Antiport. The following image illustrates the three classes of
proteins with examples of each.
Symport
A symport transports two different molecules across the membrane in the same direction in a cooperative manner.
Click the green arrow to play the animation.
Antiport
Antiports transport two different molecules through the membrane in opposite directions.
Endocytosis
While facilitated diffusion and active transport account for a great deal of the specific
uptake of molecules and ions needed by the cell, other sources of external matter can
also be taken up by the cell. The channels and pores provide a means by which molecules
can pass directly through the membrane to the cytoplasm. Other mechanisms also exist by
which molecules are taken up by the cell but do not directly pass through the plasma
membrane. This mechanism is referred to as endocytosis. This mechanism involves the
engulfing of the matter by the plasma membrane and internalization of the engulfed
material inside a cytoplasmic vesicle. Non-specific uptake of molecules occurs by
phagocytosis (large particles and macromolecules) and pinocytosis (water soluble small
molecules).
In general endocytosis, a molecule or particle encounters the cell surface and randomly
causes the membrane to create first a pit in the membrane, followed by a further
invagination of the plasma membrane and finally the pinching off of the plasma membrane
around the molecule or particle resulting in the formation of a vesicle in the cytoplasm
of the cell. Note that during the process the asymmetry of the membrane would be
apparent by the fact that the inside surface of the newly formed vesicle is the same as
the exterior surface of the cell. Thus the integrity of the cytoplasm and its exposure to only
the inside surface of the membrane is preserved. Exocytosis is just the reverse of this
process with the fusion of an internal vesicle with the plasma membrane thus releasing
its content to the outside. The balance of exocytosis and endocytosis preserves the size
of the plasma membrane and ensure neither growth nor shrinking of the cell size.
Once internalized the new vesicle fuses with a slightly acidic early endosome and
subsequently with the lysosome where the contents of the original endocytic vesicle are
digested and the digested products released to the cytoplasm where they are available
for use by the cell. This process is depicted in the following animation.
You can use the magnifier in the lower righ-hand corner to zoom the animation
Protein Transduction – an example of macropinocytosis
In the early 1990’s an intriguing observation was made that has lead to a number of new
approaches to drug delivery and therapeutic delivery systems. It is based on an
observation of the activity of the transactivator TAT protein associated with the human
immunodeficiency virus (HIV-1) and subsequently polyarginine (arginine is a positively
charge, naturally occurring amino acid) and other proteins containing a basic peptide
region referred to as the protein transduction domain (PTD). The observation was the
translocation of virtually any molecule, particle, even liposome that has the PTD
attached. While there is still some debate as to the exact mechanism of the
translocation, there is some agreement on the general process. The highly positively
charged PTD, attached to its ‘cargo’, has a tight electrostatic (ionic) interaction with
certain molecules, which are ubiquitous to all cells, in the plasma membrane. Binding to
the surface initiates macropinocytosis (pinocytosis with a slightly larger soluble
molecule). The presence of the PTD provides for high efficiency initiation of
endocytosis. While not involving a specific receptor, the binding by the PTD to the cell
surface reduces the concentration dependence for initiation of internalization.
You can use the magnifier in the lower righ-hand corner to zoom the animation
In contrast to the normal endocytic vesicle the PTD directed endocytic vesicle undergoes
retrograde transport. In this process the new vesicle fusses with the Golgi and is the
PTD containing proteins are transported back to the Rough Endoplasmic Reticulum where
they undergo post-translational modification and are transduced directly to the
cytoplasm.
Receptor Mediated Endocytosis
Much as the channels and pores discriminate between specific molecules and their
transport through the membrane, specificity and discrimination are seen during
endocytosis using specific membrane bound receptors. This targeting defines the uptake
of specific molecules or assemblies by specific cells. The following animation
demonstrates the process of receptor mediated endocytosis for the Low Density
Lipoprotein (LDL) complex. The process is divided into the individual steps to emphasize
similarities and differences among general endocytosis, protein transduction and
receptor mediated endocytosis. Several general concepts are illustrated during the
process including bioselectivity and intermediate recycling.
The plasma membrane is a very selective barrier. We have seen how some small molecule pass
freely, but most molecules are selectively brought into the cells using transporter proteins.
Most of these small molecules are metabolites or ions used in the general metabolism of the cell.
The cell also needs to transduce information across its membrane. Cells receive signals from the
surrounding fluids and other cells. These signals may tell the cell to divide or prevent division
and promote growth.
The animation below demonstrates the action of signal transduction through a G-Protein
coupled receptor. The ligand is the external signal and it binds the receptor. The G-Protein
complex is now able to bind to the receptor. This activates the G-protein by allowing the
exchange of GTP for GDP. When bound to GTP the G-protein is able to bind to Adenylate cyclase and
activate it. Adenylate cyclase generates the internal signal that is then interpreted by the cell.
Modern molecular biology is built upon our understanding of the structure and function of DNA
(deoxyribonucleic acid) and RNA (ribonucleic acid) and the enzymes and proteins that interact
with these structures. In this section of the course the structures of RNA and DNA will be
explored along with the processes by which they are used to transmit information.
The structure of DNA (shown on the left below) is the molecule upon which the Central Dogma
of modern molecular biology is based. It contains the information necessary to code for the
RNA and proteins used by a cell or virus to replicate and produce the next generation. While a
virus does not satisfy one of the major tenets of the Cell Theory that the entity is able to
self-replicate, it still uses information from either its own DNA (RNA) or its host DNA to
replicate itself.
Although RNA has much of the same basic structural features of
DNA, it takes on many more tertiary structures and has multiple
functions in the cell. The forms of RNA include:
- mRNA: messenger RNA is a copy of the DNA sequence that is read by
the ribosome during protein synthesis. The mRNA contains the
information on the order of the amino acids in protein that is being
synthesized on the ribosome. Each base triplet corresponds to one amino
acid.
- tRNA: transfer RNA is responsible for translation of the nucleic
acid code to an amino acid. tRNA carries an amino acid to the protein
synthesis machinery (ribosome) and is responsible for decoding the mRNA
information to insert the correct amino acid into the growing protein
on the ribosome. tRNA folds into a distinct three dimensional
structure, as shown in the 3D Jmol image on the right.
- rRNA: ribosomal RNA is a significant structural component of the
ribosome and plays a role in the catalyzing the formation of the peptide
bond during protein synthesis. Since the rRNA catalyzes the reaction,
it is referred to as catalytic RNA or a ribozyme.
Combining all of these roles together, the Central Dogma of modern molecular biology follows
the process outlined in this section and depicted below. DNA, with the aid of specific
proteins and enzymes is replicated (DNA Replication) thus providing a copy to be passed to the
next generation, the DNA is transcribed in sections (Transcription) into a RNA molecule that
codes for a protein or that can itself be used in the form of RNA, and the RNA can be
translated (Translation) into the primary sequence of a protein.
This short animation summarizes the processes central to Molecular Biology. The DNA
double strand separates, the RNA Polymerase moves along the strand and transcribes the
information from one strand of the DNA to a strand of RNA, and the RNA leaves the nucleus
where its sequence is translated to a protein sequence on the ribosome.
The intensive efforts to understand the structure of the molecules involved as well as the
details of the process have also yielded a set of tools that have led to the sequencing of
whole genomes containing the inherited information passed from generation to generation. This
immense amount of information has spawned the field of computational biology that is able to
extract information from the sequences that can be applied to both our basic understanding of
the functioning of organisms and to applications leading to potential cures for genetically
inherited diseases.
The Role of DNA and RNA
This section begins our description of the structure and function of DNA and RNA. The ultimate
tertiary structures of RNA and DNA are dependent on both the similarities and differences
in the primary structure of each of the polymers.
Both DNA and RNA are linear polymers of building blocks.
Each block contains a planer nucleotide base that is joined to a sugar,
either a deoxyribose in the case of DNA or a ribose in the case of RNA.
Each block is joined by a oxygen-phosphate-oxygen bridge. The
alternating ribose-phosphate-ribose is referred to as the backbone
of the nucleic acid polymer, in much the same way the N, alpha-Carbon,
and carbonyl atoms in an amino acid form the backbone of a protein.
Similarly, the nucleotide bases are analogous to the amino acid
sidechains.
Protein and Nucleic Acid Structures
 The
main features of a protein (top) and a nucleic acid (bottom) polymer
are illustrated. Both polymers contain repeating units (amino acids,
ribo-bases) linked by bonds (peptide, phosphodiester) to give linear
polymers. The mainchain atoms form the linear chain. The sidechains
project off of the mainatoms, these are either amino acid sidechains
(proteins) or bases (nucleic acids). There is also a defined direction
to the chain. In the case of proteins the free amino terminus defines
the beginning, and the sequence is simply the order of the amino acids,
named here by their sidechains (side1-side2...). In the case of nucleic
acids, the ribose has a 5' end and a 3' end. The 5' end is considered
to be the beginning and the nucleic acid sequence is given as the order
of the bases, beginning at the 5' end.
The
main features of a protein (top) and a nucleic acid (bottom) polymer
are illustrated. Both polymers contain repeating units (amino acids,
ribo-bases) linked by bonds (peptide, phosphodiester) to give linear
polymers. The mainchain atoms form the linear chain. The sidechains
project off of the mainatoms, these are either amino acid sidechains
(proteins) or bases (nucleic acids). There is also a defined direction
to the chain. In the case of proteins the free amino terminus defines
the beginning, and the sequence is simply the order of the amino acids,
named here by their sidechains (side1-side2...). In the case of nucleic
acids, the ribose has a 5' end and a 3' end. The 5' end is considered
to be the beginning and the nucleic acid sequence is given as the order
of the bases, beginning at the 5' end.
As with proteins, the bases in the nucleic acid can
interact with each other to form complex structures. The most important
type of interaction between the bases is hydrogen bonding. If bases on
two strands have complimentary hydrogen bonding properties they can
form a basepair. Double stranded DNA consists of two
anti-parallel hydrogen bonded strands. RNA can also exist as an
anti-parallel two stranded structure or it can assume more complicated
structures, such as the tRNA molecule shown on the previous page. The
detailed chemical structure of DNA and RNA will be explored in more
detail in the following pages.
The Backbone
Both DNA and RNA are linear polymers. The components of the backbone of the polymer
include a set of furanose
sugars linked together by bridging phosphate molecules in a synthesis between
the 3 position of one furanose and the 5 position of the next furanose. This linkage is
made through condensation synthesis formation of ester bonds by a bridging phosphate
molecule between two hydroxyl groups, one on each furanose ring. The resulting polymer is
a string of furanose molecules linked by phosphodiester bonds and having the 5 position
exposed on one end of the polymer and the 3 position exposed on the other end of the
molecule.
Polymer
The major difference in the polymer backbones between DNA and RNA is the sugar used in
the formation of the polymer. In DNA (DeoxyriboNucleic Acid)
the sugar is the aldose deoxyribose with a hydrogen replacing the hydroxyl at the 2
position of ribose, a furanose. In RNA (RiboNuceic Acid), the
sugar is the monosaccharide ribose in the furanose conformation.
The numbering of the positions on the sugar furanose rings of DNA and RNA follow a
convention that uses ' (prime) to denote the sugar positions. Thus the ribose has X
connected to the 1' position and hydroxyl groups on the 2', 3' and 5' positions. Using
this nomenclature, deoxyribose is formally called 2'-deoxyribose (2 prime deoxyribose) to
denote the loss of the hydroxyl at the 2' position of ribose.
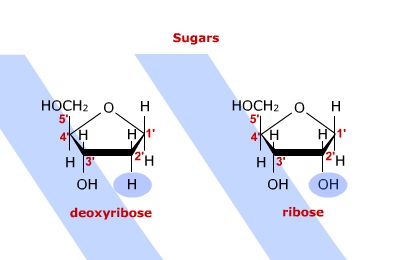 Furanose Sugars
Furanose Sugars
And, the resulting phosphodiester link is between the 3' position of one furanose and the
5' position on the second furanose.
Formation of phosphodiester Bonds
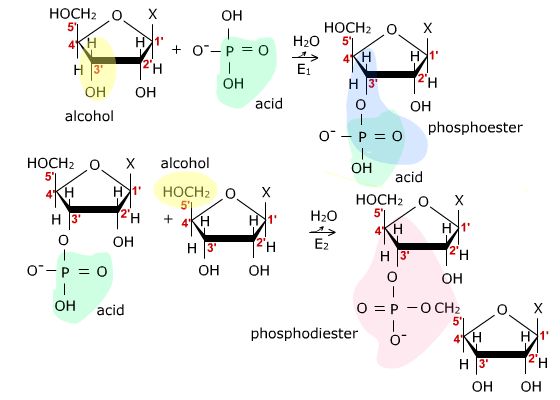 Condensation synthesis of the phosphodiester bond between the 3' position of one
ribose and the 5' position of the second ribose.
Condensation synthesis of the phosphodiester bond between the 3' position of one
ribose and the 5' position of the second ribose.
The images below show the backbone structures for both DNA and RNA.
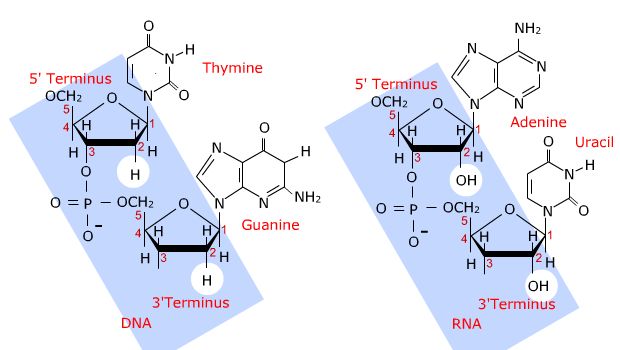 Phosphodiester linked sugar-phosphate-sugar backbone
Phosphodiester linked sugar-phosphate-sugar backbone
The synthesis of the backbones illustrated above is directed by specific enzymes that
restrict the possible structures to links between the 3' position and the 5' position of
the sugars with a bridging phosphate and to selected sugars with specific inclusion of
either ribose or 2'-deoxyribose.
The following is a list of structural characteristics of the DNA/RNA polymer backbone.
- Phosphate-ribose(deoxyribose)-phosphate-ribose(deoxyribose)sequence
- Linked by Phosphodiester covalent bonds
- 3' position on one ribose(deoxyribose) linked to 5' position of adjacent
ribose(deoxyribose) through phosphodiester bridge
- chain has 3' end and 5' end
In the structure above the X represents the bases that distinguish the units of the
backbone from each other in much the same way that the 20 naturally occuring side chains
on a common backbone distinguish the units of a polypeptide (protein) structure. These
bases will be explored in the following sections.
The following is a video in which Dr. Brown explains the structure and formation of
DNA and RNA backbones.
The Bases
The backbone structure of both DNA and RNA shows variability according to the composition
of X in the structure given on the previous page. X represents a set of nitrogenous bases.
The bases are divided into two fundamental ring structures: purines (2 fused aromatic
rings) and pyrimidines (a single aromatic ring). The differences in the bases, which are
only found attached to a sugar, give rise to the second major variation in the difference
between DNA and RNA.
All of the bases have the common structural characteristic that they are planar
structures due to the aromatic (alternating double bonds in the rings) structure of
the molecules. This feature parallels that previously seen with the phenyl group [Refer to
the Functional Groups simulation in the Glossary to review the structure of the phenyl
group]. This aromatic character also gives rise to the ability of the bases to absorb
ultraviolet light at 260 nm. This latter feature provides a distinctive means for
identifying the presence of DNA and RNA molecules.
Purine
|
Pyrimidine
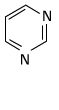
|
|
2 fused, planar rings
|
one, planar ring
|
The variations in the structures of the purine and pyrimidine bases are limited to the
five that are used during copying of DNA (Replication) or transcribing of DNA into RNA
(Transcription). The structures of these nitrogenous bases are given below.
The bases do not occur as free bases in nature but are always bound to a furanose ring.
Since the bases are always associated with a sugar in nature, they take on specific names
according to their structure. The base by itself has a specific name and the base attached
to a sugar (a nucleoside) has a distinct name. The following table gives the names of the
purines and pyrimidines as the free base and as the nucleoside with their one letter
abbreviation. The naming of the is analogous to the
with the added specification of the location of the phosphate group. An example
of this nomeclature is given below the table with the structures.
Structure of Bases
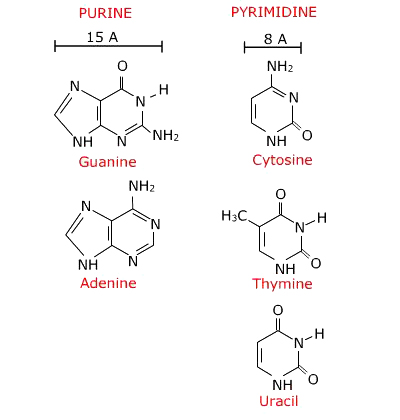 The Purine and Pyrimidine Bases
The Purine and Pyrimidine Bases
| Purine Bases |
Pyrimidine Bases |
| base |
nucleoside |
found in |
base |
nucleoside |
found in |
| Adenine |
Adenosine (A) |
DNA and RNA |
Thymine |
Thymidine (T) |
DNA only |
| Guanine |
Guanosine (G) |
DNA and RNA |
Uracil |
Uridine (U) |
RNA only |
| . |
Cytosine |
Cytidine (C) |
DNA and RNA |
Nucleoside vs Nucleotide
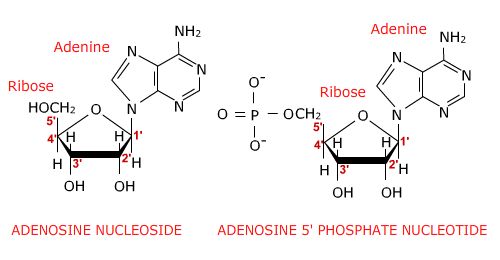 A nucleoside contains only a base attached to a sugar while a nucleotide is
composed of a nucleoside to which a phosphate has been added at either the 3' or 5'
position.
A nucleoside contains only a base attached to a sugar while a nucleotide is
composed of a nucleoside to which a phosphate has been added at either the 3' or 5'
position.
Linking the nucleotides together, a linear polymer is
generated with sequences varying
according to the placement of the bases along the final
backbone. The following figure shows a tetranucleotide sequence of DNA.
The bases, sugars and phosphodiester bonds are alternately highlighted.
In the lower left of the frame is a schematic representation of the same
sequence.
The Phosphodiester Bonds between the sugars form the backbone structures of the DNA and RNA
The illustration below shows a dinucleotide of DNA and RNA. Note
the backbone to which each of the bases is attached. While there are
four bases associated with DNA and four with RNA, are they the same set
in each case?
Dinucleotides of DNA and RNA
 Replacing the X's on the Backbone of DNA and RNA
Replacing the X's on the Backbone of DNA and RNA
The following link is a video in which Dr. Brown explains the structure of bases.
Hydrogen Bonding Between Bases
The following is an interactive simulation that allows you to form hydrogen bonding pairs
between the appropriate bases in DNA and RNA that would be allowed in the formation of the
double helical structure. You should apply the definition of the hydrogen bond to form all
possible hydrogen bonds in any pair of bases you choose. All of the possible hyrogen bonds
may be useful later as we explore multiple structures especially for RNA. The focus of
this exercise is to itentify bonding partners that will be optimal in the formation of the
DNA and RNA helical structures.
Select the proper base
from the right to bond with the example on the
left. Draw the hydrogen bonds between then and then click the
Done when you have drawn all of the bonds and then identify the bases.
Hybridization of DNA and RNA
On the previous page you determined that the most stable
complimentary base pairing takes place between A and T with two hydrogen
bonds and between G and C with three hydrogen bonds in DNA. Combining
that finding with the backbone information that described DNA as
containing A, T, G, and C as the possible bases and A, U, G, and C as
the possible bases in RNA, the complimentary base pairing in RNA would
include A with U (examine the difference in structure between T and U)
with two hydrogen bonds and G with C as described in DNA. In each case
these comlimentary base pairings included a purine hydrogen bonded to a
pyrimidine. This means that the distance between the attachment sites
for the sugar in all the base pairs are identical giving uniform
dimensions to the distance between the two backbones along the length of
two strands that are hydrogen bonded together.
Because T and U are identical in the hydrogen bonding
that each makes with A, this means that a backbone (strand) of DNA can
hydrogen bond with another strand of DNA but also with a strand of RNA.
The ability of the strands of DNA and RNA to hydrogen bond with each
other either as homodimers (DNA-DNA, RNA-RNA) or as heterodimers
(DNA-RNA) is referred to as hybridization.
As a general rule, hybridization will take place to maximize the
number of hydrogen bonds that can be formed between two polynucleotide
strands. However, another structural limitation is placed on the
formation of the hydrogen bonds between two lengths of polynucleotide
strands: the strands must be anti-parallel to each other. This means
that the since the backbne has a 5' end and a 3' end, two strands
hydridized to each other must have one strand oriented 5' to 3' and the
complimentary strand 3' to 5' as illustrated below.
| 5'- |
G- |
C- |
A- |
U- |
A- |
G- |
-3' |
| 3'- |
C- |
G- |
U- |
A- |
U- |
C- |
-5' |
This short duplex structure is anti-parallel and has 15 hydrogen
bonds holding the two strands together. As written this is a duplex of
two strands of RNA. The figure below shows the same structure as a
duplex of DNA.
| 5'- |
dG- |
dC- |
dA- |
dT- |
dA- |
dG- |
-3' |
| 3'- |
dC- |
dG- |
dT- |
dA- |
dT- |
dC- |
-5' |
Can you generate a hybrid structure using the top 5' to 3' DNA strand
from the figure above and create the appropriate RNA strand hybridized
to it?
Stability of DNA - Base Stacking and Hydrogen Bonds
The stability of double stranded DNA is due to two
factors, hydrogen bonds and base stacking. Hydrogen bonds provide an
attractive force between the strands while base stacking (van der Waals)
stabilizes the helical structure. The Jmol below shows the stacking of
the bases and illustrates the hydrogen bonding
pattern for a GC basepair. The stability of the DNA is to some extent
determined by the number of hydrogen bonds holding the two strands
together. Thus DNA with more GC base pairs is likely to be more stable
than one with more AT base pairs.
Experimentally it is possible to measure this stability by
following the melting of DNA with increasing temperature. Below is a
graph of the melting of DNA that coincides with an increase in the
ultraviolet (UV) absorbance of the DNA at 260 nm. The increase in
absorbance is referred to as the hyperchromic effect and is a measure of
the breaking of the hydrogen bonds between the bases and the separation
of the two strands.
Measurement of Melting Temperature (Tm) of DNA
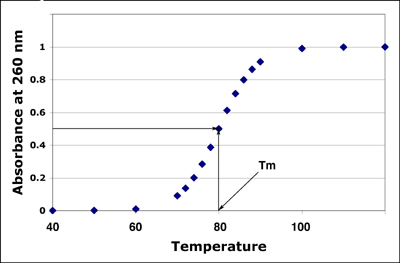 The melting temperatre (Tm) for a piece of DNA is the temperature at which 50% is no longer hybridized.
The melting temperatre (Tm) for a piece of DNA is the temperature at which 50% is no longer hybridized.
The midpoint of the transition from the double stranded to the
single stranded form of the DNA is called the melting temperature (Tm)
for the DNA. In most organisms the Tm of the chromosomal DNA ranges from
85-100 degrees C. It is possible to determine the composition of the
DNA experimentally from its Tm because the Tm of DNA is directly
proportional to the GC content of the DNA as graphically illustrated
below.
Measuring the composition of DNA
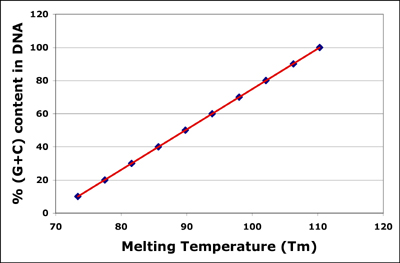 The
melting temperature is directly proportional to the composition of the
DNA. This is also a measure of the hydrogen bonding content of the DNA.
The
melting temperature is directly proportional to the composition of the
DNA. This is also a measure of the hydrogen bonding content of the DNA.
While the melting temperature does not tell us anything about the
proteins that are coded by the DNA, it does tell us something about the
tolerance of the organism in which the DNA is found. For example,
thermophilic bacteria (those that survive at extremely high
temperatures) have DNA with very high Tm values so that their DNA does
not melt at their normal environmental temperature. As we will see
later, this also has implications for how the same proteins can be made
in two different organisms using DNA with vastly different compositions.
In summary, for the hybridization of two strands of DNA and RNA
- the chains are antiparallel
- the two chains are held together by hydrogen bonding between bases
- the stability of the DNA is directly proportional to the number of hydrogen bonds between chains
- hydrogen bonding between strands of DNA and RNA follow the pattern
- A hydrogen bonded through two bonds to T in DNA
- A hydrogen bonded through two bonds to U in RNA
- G hydrogen bonded through three bonds to C in both DNA and RNA
Building the Double Helix
As described in the previous section, the hybridization of DNA and RNA results in the formation of:
- Double stranded DNA or RNA with
- Antiparallel orientation (5’ to 3’ against 3’ to 5’) with
- Uniform distance between the strands due to pairing of a purine with a pyrimidine (A with T (or U) and G with C).
This representation of the ladder of a double stranded DNA is
illustrated in the figure below on the left side. However, this
secondary structure spontaneously forms a double helical coiled
structure that is depicted on the right hand side of the figure below.
This structure has several distinct features that characterize the
dominant structure called B-DNA.
Use the interactive 3D Jmol image to explore these features of the double helix:.
- The two helical polynucleotide chains are coiled around a common axis
- The phosphate and ribose groups are on the outside
- The bases are directed toward the inside of the helix, and stack on the central helical axis.
- The planes of the bases are perpendicular to the axis of the helix
- The diameter of the helix is uniform and is 20 Angstroms
- The bases are stacked and separated by a uniform distance of 3.4 Angstroms.
The following link is a video in which Dr. Brown explains how DNA and RNA are constructed from the backbones and bases.
DNA has Grooves
A predominate feature of the DNA double helix is the presence
of grooves, or indentations, on the side of the helix. This grooves
expose the edges of the basepairs, The grooves are called the major
groove and the minor groove. Both grooves are deep and expose the edges
of the bases to the external environment, making them accessible for
protein binding. The minor groove is quite narrow (approximately 12
Angstroms across) and while the edges of the bases may be accessible to
solvent and small molecules, they are generally not accessible to larger
molecules. The major groove on the other hand is quite wide
(approximately 22 Angstroms across) and is sufficiently wide to
accommodate a protein alpha helix. Below is a side view of the B-DNA
structure using a space filled model (left) next to a 3D Jmol image that
will allow you to highlight each groove and determine which atoms from
each basepair project into the groove. The nature of the atoms that
are exposed in the grooves of the structure are important for the
ability of proteins to recognize specific sequences of DNA.
Discrimination of the different sequences must be made by having access
to the bases inside the structure since the backbone structure is common
to all sequences of DNA
Side View of B-DNA Helix
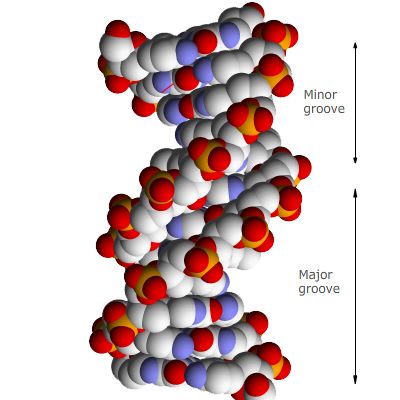 This space filled model of the B-DNA shows the major and minor grooves resulting from the formation of the double helix. This space filled model of the B-DNA shows the major and minor grooves resulting from the formation of the double helix.
|
|
|
Bases pairs inside double helix
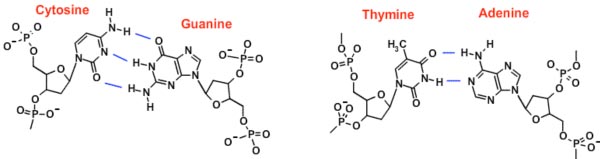 Use the buttons on the above Jmol to see the location of a C-G and a T-A basepair in the double helix.
Use the buttons on the above Jmol to see the location of a C-G and a T-A basepair in the double helix.
The tertiary structure of B-DNA has been described with a number of specific characteristics.
These characteristics describe what is referred to as the B-helix. It is the most common
structural form of double stranded DNA. RNA and hybrids of DNA with RNA have altered
characteristics resulting from the substitution of a ribose for a deoxyribose in the backbone
structure. The general characteristics found in the B-DNA are still present; however, slight
differences in the specific characteristics result in dramatic changes in the overall
structure of the helix.
The following table compares some of the finer characteristics of the B-helix and A-helix.
In general, the A-helix is broader than the B-helix, there are slightly more base pairs per
turn in the A-helix, and most significantly, the tilt of the bases relative to the axis of the
helix is much greater in the A-Helix.
| |
A-helix |
B-helix |
| Shape |
Broader than B |
|
| Screw Sense |
Right |
Right |
| Base pairs/Turn |
11 |
10.4 |
| Pitch/turn |
25 |
35 |
| Tilt of Bases |
19 |
1 |
| Major Groove |
Narrow/Very Deep |
Wide/Quite Deep |
| Minor Groove |
Very Broad/Shallow |
Narrow/Deep |
The following 3D representations illustrate the result of these
differences in comparing similar views
of the B-helix of DNA with the A-helix in duplex RNA. When
exploring these structures you should focus on the following aspects of
the structures, noting similarities and differences:
- The location of phosphate, ribose, and bases with respect to the interior and exterior of the helix
- The location of the major and minor grooves. The grooves are water
and ion filled channels on the side of the double helix. Proteins
usually interact with the edges of the bases in the major groove.
- The tilt of the bases with respect to the axis of the helix.
- How the bases overlap with each other. As a result of the greater
tilt in the bases relative to the helix axis, the bases of the
A-helix slide past each other and do not exactly stack on top of
each other as they do in the B-helix.
As discussed at the beginning of this unit, modern molecular biology has developed a Central
Dogma that describes a series of processes starting with DNA and ending with the production of
a protein using the genetic code. This Central Dogma is diagrammed in the figure below. The
initiation of this process is DNA Replication, highlighted in red, which describes the copying
of the information existing in DNA to new DNA. One of the most important processes that a cell
performs before it can divide is to faithfully replicate its chromosome. While there are
differences between prokaryotes
and eukaryotes, there are great similarities in the mechanism by which they replicate their DNA.
This unit will describe this mechanism of DNA replication
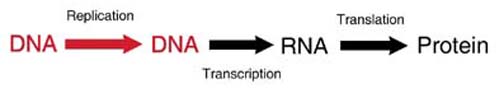
In addition, an understanding of the enzymes that are involved in faithfully copying the DNA
has also lead to their use in applications that have advanced our knowledge of genomes, the
information they contain and how they are used. The polymerase chain reaction (PCR) and DNA
sequencing using dideoxynucleotides have revolutionized our ability to work with small amounts
of DNA and generate immense amounts of sequence information in a very short time. The
understanding of the basic principles involved in DNA replication have also lead to our
understanding of such topics as DNA repair and chromosome extension.
DNA replication is the process by which DNA is copied resulting in a faithful replicate
of the original double stranded DNA. Key to the inheritance of the code for materials
produced and used in a cell, DNA replication must be faithful thus not allowing or
minimizing errors during the process and it must generate new DNA that can be transmitted
to the next generation. Because DNA replication generates a copy of the original, it is
referred to as DNA dependent DNA synthesis. It is dependent on DNA because it
must have a template to use in making the copy and the result is a new strand of double
stranded (duplex) DNA.
Semi-conservative Replication
The process of DNA replication requires that the original DNA act as a template
for the newly formed DNA. How the new DNA acts as a template could occur in multiple ways.
As depicted below the double stranded DNA (anti-parallel strands indicated by the grey and
opposite polarity blue strand) could be completely copied leaving the original strands of
DNA intact (grey) and yielding a newly synthesized double strand DNA containing two new
strands (blue). This is referred to as conservative replication. The second model of DNA
replication starts with the same double stranded DNA but the products each contain one of
the original strands and a complementary newly synthesized strand. This is referred to as
semi-conservative replication. Biological organisms have been shown to
replicate their DNA by semi-conservative synthesis.
The “Learn More…” that follows shows the Meselson-Stahl experiment that was performed to
prove that semi-conservative replication is the mechanism within biological systems.
Semi-conservative vs Conservative replication models

DNA synthesis
DNA synthesis in biological systems is a very directed process controlled by specific
enzymes that both dictate where synthesis begins and the exact structural conditions
required to undertake DNA synthesis. Knowledge of these conditions and the process
involved in DNA synthesis has become the basis for understanding other biological
processes involving DNA and in the design of a set of tools essential to the advancement
of the study of modern molecular biology.
DNA polymerases (setting the rules)
DNA polymerase is an enzyme that has very specific substrate requirements and
thus controls where the synthesis of DNA can take place. An understanding of the basic
mechanism of this enzyme will make the understanding of how DNA replication works become
clear.
- As with any enzyme, DNA polymerase has a very specific set of requirements for
recognition of its substrate. For DNA polymerase the substrate is single stranded
DNA not double stranded DNA. Thus DNA polymerase is not able to bind to or
recognize an intact genome but
rather it uses each single strand of DNA as a template.
- While DNA polymerase requires single stranded DNA as a template, it does not have the
ability to start anywhere on the strand and initiate de novo synthesis. The active site of the polymerase
requires a segment of DNA or RNA bound in a complimentary manner to the single stranded
template as an initiation point for the synthesis reaction. This segment of DNA or RNA
is generally referred to as the primer.
- If you examine the structure of either DNA or RNA there are two places that a new
deoxy-nucleotide can be added to extend the length of the complementary strand.
While DNA synthesis could be from either end of the primer, during replication using the
enzyme DNA polymerase the synthesis is restricted by the enzyme to growth in only one
direction. Deoxy-nucleotides are only added to the 3’ end of the primer and thus
synthesis is unidirection from 5’ to 3’.
- For synthesis to proceed, the enzyme imposes a third restriction on the substrate. Not
only does it require a DNA template and a primer, but the primer must have a free,
3’ hydroxyl group for the addition of the next deoxy-nucleotide.
- And finally, all four deoxy-nucleotide triphosphates must be present to completely
synthesis the complementary strand of DNA. For DNA synthesis (replication) this means
dATP, dTTP, dCTP, and dGTP must be present. Collectively these are referred to as dNTPs
(deoxy-Nucleotide TriPhosphates).
More than one DNA polymerase enzyme is known and each carries out a specific function
depending on the situation. For example, DNA polymerase III is the enzyme used in
replicating a genome while DNA polymerase I is used in DNA repair. Even with clear
differences as to when each of the DNA polymerases is used during the replication of DNA,
the basic substrate requirements are the same.
The following diagram depicts the structural requirements for the operation of DNA
polymerase enzymes.
Perfect Substrate for DNA Polymerase
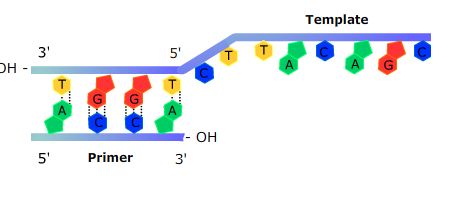
Synthesis (the making of new DNA)
Once the appropriate structural requirements for the synthesis of DNA are met, DNA
synthesis occurs continuously in a unidirection process from 5’ to 3’ along the template.
The following animation demonstrates the building of a complementary strand using the
components required by DNA polymerase. This process can take place wherever the initial
conditions for operation of DNA polymerase exist.
DNA Polymerase binds to DNA at the 3' end of the primer and builds a
complementary sequence.
Initiation of synthesis from double stranded DNA
Having explored the requirements for DNA polymerase, it is clear that the prokaryotic and
eukaryotic genomes fail the first requirement of a single stranded sequence of DNA.
Therefore to initiate DNA replication it is necessary to create the conditions that will
allow DNA polymerase to carry out DNA synthesis.
- First the DNA must be opened by breaking the hydrogen bonds between the base pairs and
unwinding the DNA. This is initiated by a protein called the initiator protein or
DnaA that recognizes a specific DNA sequence called the origin of
replication, binds to the sequence and opens up the sequence to create a
replication bubble (open complex). An enzyme called a helicase binds
to the open complex and extends the melted region of the double stranded DNA to further
open the replication bubble. There now exists a single stranded segment of DNA exposed
on both chains of the DNA double helix. A small protein called SSB (Single
Stranded Binding protein) coats the open complex.
The Open Complex
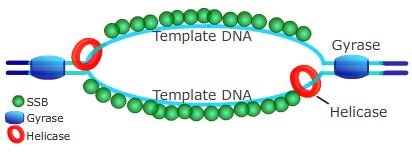
- With formation of an open complex, one of the requirements for DNA polymerase is
satisfied: single stranded DNA is generated. At either end of the open complex where the
single stranded and double stranded DNA meet, a structure referred to as the
replication fork exists. From this point we will focus on the process of
replication at one of the replication forks. Remember that the replication will be occurring
simultaneously from both forks in opposite directions. To allow the binding of DNA
polymerase, a second requirement must be met for the structure of the substrate. An enzyme called Primase synthesizes a short
(approximately 10 nucleotides), complementary RNA primer on each strand of the DNA in a
5’ to 3’ unidirectional fashion. This complex of DNA and protein at the replication fork is referred to as the replisome.
Replisome at one fork

- The substrate requirements for DNA polymerase have been satisfied and synthesis
begins. Helicase continues to melt the DNA in front of DNA polymerase.
- As single stranded DNA is generated between the ‘back’ of the RNA primer on one
strand and the opening replication fork, a new segment of RNA primer is laid down near
the opening fork and DNA polymerase synthesizes a new complementary strand of DNA from
the primer toward 5’ end of the existing primer and stops. This segment of DNA that is
not connected to the next segment is referred to as an Okazaki Fragment. Synthesis on
this lagging strand is said to be discontinuous since it is generated in uniform lengths
of DNA starting from one primer and stopping before the next RNA primer.
- Synthesis on the opposite strand is a continuous uninterrupted process starting
from the initial primer and continuing as long as the DNA continues to open up at the
replication fork. This strand on which continuous synthesis takes place is referred to
as the Leading Strand.
 Okazaki fragments on the lagging strand
Okazaki fragments on the lagging strand
The following animation depicts the complete process of DNA replication. In the animation
you should be able to identify each of the stages of synthesis on each of the strands. At
the end of the synthesis you will see that the leading strand has a continuous double
helix generated while the lagging strand has a discontinuous set of Okazaki fragments that
must be connected before the synthesis is complete.
To accomplish this mending of the DNA, a separate DNA polymerase binds to the ‘nick’ left
in the DNA between the 3’ end of the newly synthesized DNA and the 5’ end of the RNA
primer. The substrate requirements for DNA polymerase are met even though there is no
apparent single stranded DNA present. The DNA polymerase removes the RNA primer and a few
nucleotides of DNA in front of it while simultaneously synthesizing DNA on the 3’ end of
the existing DNA. This process is referred to as nick translation.
When only DNA is present, the DNA polymerase releases from the DNA leaving a nick in the
DNA between two DNA segments. DNA polymerase does not have a function to join the two ends
of the DNA together and thus a separate enzyme, DNA Ligase, recognizes the nick in the DNA
and links the 3’ end of the one segment to the 5’ end of the next segment.
This process of DNA replication continues until the entire sequence of DNA is
synthesized. The process is fundamentally the same in prokaryotes and eukaryotes.
Solving Challenges within the Rules
The Origin of Replication – where to start
The Challenge: To initiate the DNA replication of a chromosome the appropriate
substrate conditions must be met as described previously. How long the process of
replication takes is also important to the cell. How genomes of variable sizes can be
totally replicated within the appropriate life cycle time of the cell is a significant
challenge.
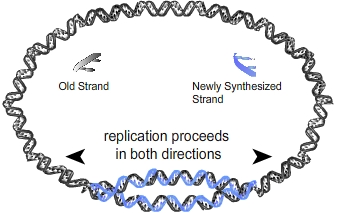 Bacterial chromosomes have a single point of origin for replication
Bacterial chromosomes have a single point of origin for replication
The origin of replication has been described as a specific DNA sequence to
which initiator protein (DnaA) binds and initiates the process of creating the open
complex for synthesis. For a typical prokaryotic (bacterial) chromosome there is one
origin of replication and synthesis proceeds at a rate of approximately 500 bp(base
pairs)/sec. A bp or base pair is the unit of length of DNA measured in single
nucleotides or hydrogen bonded base pairs along the polymer chain. Thus if a typical
bacterial chromosome is 2x10e6 (two million) bp in length then the replication time of the
chromosome would be about 30 minutes which is consistent with the life cycle of the
typical bacterial being 30 to 40 minutes.
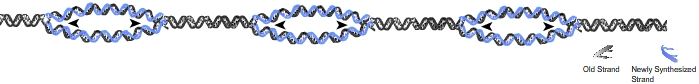 Eukaryotic chromosomes have multiple points of origin for replication
Eukaryotic chromosomes have multiple points of origin for replication
The replication rate for the eukaryotic replication fork has been measured to be
approximately 50 bp/sec. This means that for a eukaryotic chromosome of 10e8 bp to
completely replicate starting at one origin of replication it would take approximately 23
days. While we know that the life-cycles of eukaryotic cells are extremely variable, the
S-phase, the period of time during the eukaryotic life-cycle when DNA is replicated,
typically last 8 hours. To explain the enormous difference in time scale, it has been
found that the eukaryotic chromosome has multiple origins of replication. The
eukaryotic replication units have upwards of 80 origins which means the replication time
is reduced to approximately 7 hrs.
What if DNA polymerase makes a mistake
The Challenge: DNA polymerase synthesizes DNA at rates up to 500 bp/sec. There
are basically three types of mistakes that the enzyme can make during the synthesis:
substitution of an incorrect base, insertion of an extra base or failure to insert a base
resulting in a deletion. The consequences of these errors will be apparent as we move
toward the discussion of DNA transcription and RNA Translation. In the context of
Replication it is important to touch on at least a couple of the means by which the system
corrects errors or reduces the frequency of errors.
Editing the Synthesis
DNA polymerases are members of a very interesting family of enzymes because of the multi
functional nature of their activity. As has been discussed previously, they are capable of
synthesizing DNA given the appropriate substrate. They have their own activity to remove
RNA and/or DNA in front of the synthesis as described on the lagging strand of the
replication fork. Equally important to the organism is the ability to edit errors during
synthesis by either preventing the incorporation of the incorrect base and/or by removing
an incorrectly incorporated base before the next base is added. These editing functions
can reduce the error frequency by 2 orders of magnitude.
However, even this editing function does not get the error rate to an acceptable number
for survival of an organism. In addition to editing during synthesis, there is also
monitoring of the DNA for missed errors resulting from replication and for
post-replication damage to the DNA. In each of these cases the process involves cutting
one of the strands of the DNA specifically creating a condition by which a DNA polymerase
can replace the error or damaged DNA followed by ligation of the nicked DNA using DNA
ligase. It is to be noted that while different proteins may be used, the basic process is
the same and follows the direction/rules dictated by the mechanism of the specific
enzymes. The combination of these other editing and repair functions reduces the error
rate by another 2 orders of magnitude thus reducing the error frequency to an acceptable
level.
Following the rules
Understanding the process/mechanism of DNA replication has allowed researchers to
understand how certain DNA repair functions are conducted and to develop new tools that
have revolutionized our ability to gain information from DNA.
Overcoming Chromosome Shortening
During replication of circular chromosomes, such as in bacteria, the leading strand of
synthesis from one replication fork eventually meets the lagging strand of synthesis for
the other replication fork and there is complete synthesis of the entire chromosome. For
linear chromosomes this same process happens between two origins of replication. However
at the end of the linear chromosomes, the telomeres, the lagging strand does not
completely replicate to match the length of DNA generated on the leading strand. Thus for
repeated replication of the chromosomes during multiple cell divisions, this means that
the chromosomes are shortened with each cycle of replication.
Since DNA synthesis is always unidirectional from 5’ to 3’, it is necessary to lengthen
the 3’ end of the chromosome such that primer directed synthesis can take place to
lengthen the shorter 5’ end of the chromosome. While the staggered end of the chromosome
is not avoided, the net length of the chromosome is recovered or even lengthened. The
diagrams below demonstrate how lengthening of the chromosome can be accomplished using the
enzyme telomerase. The telomerase enzyme is a polymerase like enzyme that carries
its own template with the template having overlap complementarity with the 3’ end of the
chromosome. The criteria for polymerization are met, a template and a primer with a free
3’ hydroxyl, and the telomerase lengthens the 3’ strand of the chromosome. This process of
lengthening is repeated until a the normal replication process lays down a RNA primer
using the primase enzyme, DNA polymerase synthesizes new DNA from the 3’ end of the RNA
primer and DNA ligase joins the newly synthesized DNA to the 5’ end of the chromosome.
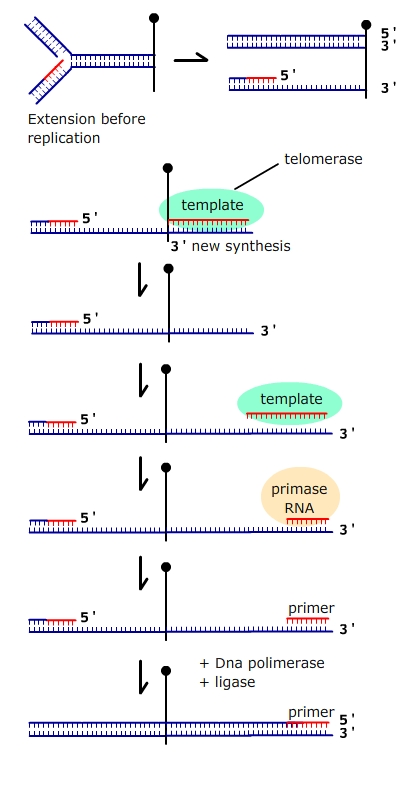 Process of Telomerase lengthening the 3’ strand of the chromosome.
Process of Telomerase lengthening the 3’ strand of the chromosome.
Being creative with the rules
Two tools that have become integral parts of the study of modern biology were developed
through the simple understanding of how the DNA replication process works. At the
beginning of this unit three structural requirements for the substrate of DNA polymerase
were described. Satisfying those requirements is the basis for the polymerase chain
reaction (PCR), used to amplify segments of DNA, and the Sanger DNA
sequencing method used in sequencing the human genome as well as others.
Polymerase Chain Reaction
Characterization of large numbers of microorganisms, rapid screening for genetic diseases
and identification of individuals or species from very small amounts of tissues or cells
requires a means to amplify a specific sequence of DNA for further manipulation.
Previously this would have required growing large quantities of cells containing the DNA
or obtaining large quantities of the tissue. DNA replication is the process of doubling
the original amount of DNA. The polymerase chain reaction (PCR) is simply a method by
which DNA replication is repeated through many cycles. In the same way that doubling a
bacterial cell through 20 cycles will generate a million copies of the bacteria,
replication of DNA through 20 cycles will amplify the amount of DNA by a million fold.
 View this PCR tutorial for a quick overview of the process. (©Sumanas,
Inc.)
View this PCR tutorial for a quick overview of the process. (©Sumanas,
Inc.)
View PCR tutorial
PCR is limited by the length of DNA that can be repeatedly replicated by DNA polymerase.
Different polymerases and different conditions are described to amplify DNA of varying
lengths. Routinely, segments of DNA from 500 to 1000 bp in length are easily replicated.
The segment of DNA to be replicated is defined by the position of the primers that define
the start of the replication on each strand. These are referred to as the forward
primer and the reverse primer. The double stranded DNA to be amplified is
melted by raising the temperature high enough to fully melt the DNA creating single
stranded DNA. The sample is cooled in the presence of the primers and each primer binds to
its complementary strand. The appropriate substrate conditions for DNA polymerase binding
and synthesis are now in place. DNA polymerase is added with the dNTPs and synthesis of
DNA proceeds for as long as the reaction is allowed to progress before either the time of
the reaction ends or the DNA polymerase falls off the substrate.
The temperature is raised again to melt the DNA and create a single stranded template.
Because DNA polymerase will generally denature and irreversibly inactivate at these
elevated temperatures it is necessary to add fresh DNA polymerase at each cycle when the
temperature is lowered during primer attachment. To avoid repetitive addition of a very
expensive enzyme after each cycle and to make automation of the process feasible, there
was an advantage to finding a DNA polymerase that survives high temperatures and in fact
is active at high temperatures. Such an enzyme was found from thermophilic bacteria
(bacteria that thrive at extremely high temperatures). One such enzyme, taq DNA polymerase
from thermos aquaticus, is optimally active around 75 degrees and can withstand
temperatures as high as 100 degrees without denaturing. Thus the PCR reaction can be
performed in a closed tube with DNA to be amplified, taq DNA polymerase, excess dNTPs, and
excess amounts of the forward and reverse primers. The reaction tube is put in an
instrument called a thermocycler that can be programmed to automatically raise and lower
the reaction temperature. Now the temperature is raised to form single stranded DNA,
lowered to bind the primers and DNA polymerase with the correct substrate again replicates
the DNA. After 20 cycles of this process with each cycle doubling the number of duplex DNA
strands present, there will be 1, 048,576 copies of the DNA present.
DNA Sequencing
Current methods of DNA sequencing rely on understanding the substrate requirements of DNA
polymerase and the concept of competition. To sequence DNA as with PCR it is necessary to
create single stranded DNA that will act as the template. It is generally required that to
ensure valid DNA sequence, a double stranded segment of DNA should be sequenced in both
directions using primers at each of sequence to be defined. These primers are the
equivalent of the forward and reverse primers of PCR. However, where PCR replicates both
strands simultaneously, DNA sequencing is performed on each strand individually. Starting
with the single stranded template and the bound primer with a free 3’ hydroxyl, the
conditions are set for DNA polymerase to replicate the DNA.
If all four dNTPs were added at this point simple replication of the strand of DNA would
occur and no information would be gained. However, if one of the dNTPs was substituted by
a NTP that had no hydroxyl on either the 2’ or the 3’ position, a dideoxyNTP
(ddNTP), it would be incorporated into the growing DNA strand through the 5’ position
in a complementary position opposite its complementary base on the template strand but
synthesis from that point forward would be terminated due to the absence of a hydroxyl on
the 3’ position. The requirement for synthesis by DNA polymerase has been lost. The
dideoxyNTP is referred to as a suicide substrate.
If complete replacement of dATP with ddATP were made, then the first time a T were
encountered in the template strand, DNA polymerase would incorporate ddATP and all
synthesis would halt. If however, a mixture of dATP and ddATP were present, then when the
first T were encountered in the template, the dATP and ddATP would compete for the active
site of DNA polymerase. If dATP is incorporated, synthesis continues to the next
nucleotide and if ddTAP is incorporated synthesis halts for that one strand of DNA. Thus
in a population of template strands, a fragment of DNA starting at the primer and ending
at each T would be generated with the length of the fragment in nucleotide units defines
the location of the T in the template strand. If this experiment is performed for each
individual nucleotide, the complete sequence of the template can be accomplished.
The second phase of the Central Dogma is the process of transcription, which is summarized as DNA directed RNA synthesis.
This unit will describe the process of DNA transcription and the
different categories of products and how each is processed. The
discussion will be organized around:
- The organization of the gene expression system into Operons,
- The initiation of transcription at special sequences called Promoters,
- The unidirectional RNA sythesis using RNA polymerase and how it differs from replication, and
- The post-transcriptional processing of the end products of transcription
Transcription
The process of DNA transcription is described as DNA-directed RNA synthesis.
This process, which takes place in the nucleus of eukaryotic cells, has
many similarities with DNA replication. The enzyme to catalyze the
process is RNA Polymerase that likewise has some
characteristics similar to those of DNA polymerase used in replication
but it also has significant differences.
- The DNA must be unwound starting at a specific DNA sequence
-
Replication – Origin of Replication
-
Transcription – The Promoter
- The process requires a template that is DNA
-
Replication – Complementary copies of both strands are created simultaneously
-
Transcription – A complementary copy is made of one strand
- The process does not require a primer
-
Replication – A primer with a 3’ hydroxyl is required for DNA polymerase
-
Transcription – No primer is required for RNA polymerase
- Synthesis is unidirectional in the 5’ to 3’ direction
- Synthesis requires four nucleotide triphosphates
-
Replication – deoxyribonucleotide triphosphates include dATP, dGTP, dCTP,
dTTP
-
Transcription – ribonucleotide triphosphates include ATP, GTP, CTP, UTP
The Operon
The organization of the genetic information on the chromosomes of
prokaryotes and eukaryotes is different. In each case, as indicated
below, the sequence for initiation of transcription is the Promoter and
the sequence signaling the end of transcription is the Terminator.
However, in eukaryotes there is generally one gene that codes for one
product under the control of a single promoter. In contrast, prokaryotes
generally have multiple genes, each coding for a separate product,
under the control of a single promoter. This unit of a promoter, a
terminator and the intervening gene or genes is called an Operon.
The operon also contains the controlling elements for the operon. The
control of expression of an operon is the topic for a separate unit.
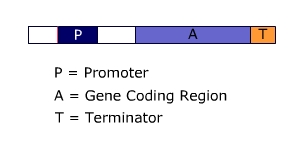 Eukaryotic Operon Eukaryotic Operon
|
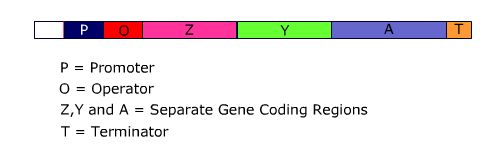 Prokaryotic Operon Prokaryotic Operon
|
The Promoter
RNA polymerase is an example of a quaternary structure composed
of a core protein for the synthesis the complementary RNA strand and a
subunit, the Sigma subunit, that first binds to the promoter region of
the Operon and creates an open complex by unwinding the double stranded
DNA. The promoter region of the operon is critical in defining how much
or the frequency with which transcription of an operon takes place.
Promoters have different sequences. The sigma subunit binds to the
different promoter sequences with different affinities. In the
illustration below, the sigma subunit binds to both of the promoters and
a dissociation constant, Kd, can be written for each binding.
 Different promoters result in different Kd values.
Different promoters result in different Kd values.
Promoters can be characterized as being strong or weak promoters
depending on their affinity for RNA polymerase. The result of a strong
promoter with a high affinity (tight binding) to RNA polymerase is a
greater frequency of starting transcription from that promoter.
Therefore, more product from transcription initiated at a strong
promoter is possible compared to product formation initiated from a weak
promoter in the same amount of time.
RNA Synthesis
Once RNA polymerase binds to the promoter, initiation of RNA synthesis or Transcription
proceeds. The following animation depicts the steps of transcription.
The Products of Transcription
The product of the process of transcription is RNA. There are
three distinctly different RNA products that result from transcription
based on their function within the cell.
-
Ribosomal-RNA (rRNA) is a large defined length of RNA that when
processed is folded into a tertiary structure that is the scaffolding
for the structure of the ribosome. The ribosome is a protein-RNA complex
quaternary structure where protein synthesis (translation) takes place.
-
Messenger-RNA (mRNA) is a variable length of RNA whose length
is dependent on the length of the coding needed to produce a protein by
translation. This RNA is the message that carries the information that
codes for the synthesis of a protein. It has a relatively short lifetime
in the cell.
-
Transfer-RNA (tRNA) is a set of short RNA molecules generally
75-80 nucleotides in length. Each tRNA molecule is specific for
transporting a specific amino acid to the ribosome during protein
synthesis (translation). All of the tRNA molecules have a common,
compact tertiary structure.
Each of the RNA classes (rRNA, mRNA, and tRNA) is produced during transcription using a different RNA polymerase.
- rRNA – RNA polymerase I
- mRNA – RNA polymerase II
- tRNA – RNA polymerase III
Bioselectivity describes the discrimination of a specific RNA
polymerase for a specific promoter yielding different RNA products.
Equilibrium binding of the polymerase with the promoter explains the
differential production of products using the same polymerase and
different promoter sequences.
Post-Transcriptional Modification
The products of transcription, rRNA, tRNA and mRNA, are each used
for different functions within the cell but are all essential to
protein synthesis. They each also undergo change or modification before
they are used to carry out their specific function. This process of
change after transcription from DNA to RNA is called post-transcriptional modification.
For ribosomal-RNA (rRNA)the modification involves cutting of
the original long segment of RNA transcript into fragments. The
processing of the RNA takes place in the nucleus and the fragments are
used in the nucleoli where ribosomes are assembled.
For transfer-RNA (tRNA), the RNA transcript has a
segment of the 5’ end removed, a trinucleotide that is common to all
tRNAs (ACC) added to the 3’ end, and a short segment of RNA spliced out
of the middle of the RNA sequence. In addition, several of the bases in
the structure are chemically modified after the final sequence is
generated.
The most extensively modified transcript is messenger-RNA (mRNA) in eukaryotic organisms. For this reason, the primary transcript or mRNA before it is modified is referred to as pre-mRNA.
- After transcription a “5’cap” is added to the transcript.
This “cap” is a GTP that is added through the 5’ position of the GTP to
the 5’ end of the transcript. This results in a free 3’OH of the guanine
nucleotide on the 5’ end of the transcript. As we will see in the next
unit, the 5’-cap is a signal used in binding the eukaryotic message to
the ribosome to initiate protein synthesis.
- A second post-transcriptional modification is the addition of a “polyA tail”
onto the 3’ end of the transcript. A specific enzyme adds 100 to 500
adenine nucleotides to the 3’ end. Once the mRNA is transported to the
cytoplasm from the nucleus, it is thought that the polyA tail acts as a
clock to measure the lifetime of the message. A 3’- exonuclease
is an enzyme that removes nucleotides one at a time from the 3’ end. If
one assumes that it removes nucleotides at a constant rate, the message
can be actively used from 3 minutes to 4-5 hours, depending on the
length of the tail, before it becomes inactive due to the loss of the
“tail.”
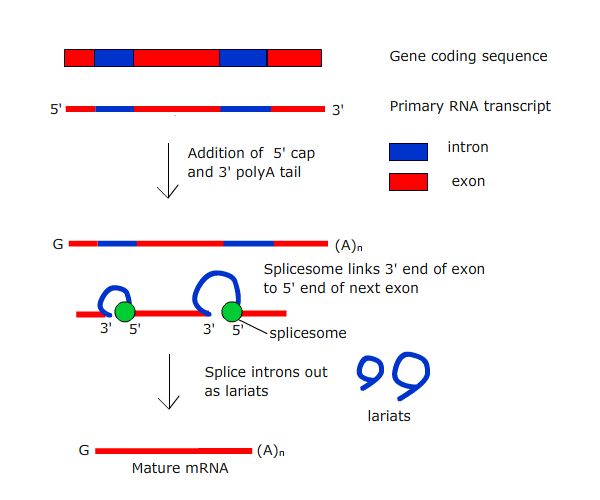
- The addition of the 5’-cap and the polyA tail are depicted in
the figure above. But, one of the most intriguing post-transcriptional
modifications is the splicing out of segments of the mRNA called Introns.
Introns are sequences that do not code for protein, are interspersed in
the original DNA and are removed from the RNA transcript by the process
of splicing. The splicing is carried out by a complex (quaternary
structure) of proteins referred to as the splicesome composed of SnRNPs (Small Nuclear RiboNucleoProteins). The segments of RNA that do code for protein are called Exons.
Signaled by nucleotide sequences in the ends of the introns, the
splicesome brings together the 3’ end of one exon with the 5’ end of the
next exon and cuts out the intervening intron. The product is the
end-to-end spliced exons and a segment of removed RNA that was the
intron in the shape of a lariat. The figure above illustrates the process. After completion of the splicing, the mature mRNA is exported to the cytoplasm where it is used in protein synthesis (Translation). Introns do not exist in prokaryotes.
Recently, research has found that a number of mRNA sequences undergo what is referred to as alternative-splicing.
If normal splicing is from the 3’ end of one exon and the 5’ end of the
immediately following exon, imagine the number of possible splices that
could occur if the 3’ end of the exon could be joined to the 5’ end of
any exon downstream from the first exon. For example, if an mRNA had
three exons, 1, 2, and 3 in order, and two introns, the normal sequence
of splicing would link exon 1 to exon 2 to exon 3. But it would be
possible to link exon 1 to exon 3 to create a “new” protein with only
two of the three exons. This allows a single gene with multiple exons to
represent many different proteins. Errors in splicing have also been
found to be the cause of certain diseases.
The final stage of the Central Dogma is translation, which is defined as RNA Directed Protein
Synthesis. This process is unique in that it takes information from one type of polymer (RNA)
and translates that linear information into the linear sequence of a totally different
polymer, a protein. This unit will describe that process and the elements that are required to
have it function. The discussion will be organized around,
- the Genetic Code and the use of the triplet code
- the role of transfer-RNA, messenger-RNA and the ribosome in translation
- the formation of the initiation complex, the elongation of the peptide, and the
termination of synthesis, and
- the consequences of errors and presentation of mutations
The Code
Translation involves the conversion of a sequence of RNA to a corresponding sequence of
amino acids. To perform the conversion a code is needed to translate from the four
nucleotides (AUGC) of mRNA to the 20 naturally occurring amino acids. The question then
is what combination of nucleotides can code for at least twenty different amino acids?
Four nucleotides taken two at a time would generate 16 possible unique sequences. Not
enough to code for 20 different amino acids. However, four nucleotides taken 3 at a time
would generate 64 different unique sequences. Clearly this is enough to code for the 20
naturally-occurring amino acids. In fact, the abundance of codes means that there is
redundancy or degeneracy in the code with more than one triplet code representing a
single amino acid.
Through experimentation it was found that the universal start code or codon is AUG and
three stop codons (UAG, UGA and UAA) were identified. The remaining 60 codons
represented the 20 amino acids. Having elucidated the entire code it was found that for
many of the amino acids, the first two nucleotides in the sequence defined the amino
acid with the third nucleotide being any of the four nucleotides. For example the codon
for the amino acid leucine is CUX where the X represents any one of the four
nucleotides, and the codon for the amino acid valine is GUX. This third base then became
known as the wobble base signifying the flexibility the system has in identifying the
third base. This also demonstrates the fact that a single amino acid can be coded by
several triplet codes but a triplet code only represents a single amino acid. This
triplet code for converting the sequence of mRNA to a sequence of amino acids is
referred to as the Genetic Code.
The Starting Materials
Having identified a code to perform the translation, it is now necessary to describe the
process by which the genetic code is used. Four major ingredients are necessary to carry
out translation: m-RNA, t-RNA, the ribosome and the initiation factors.
Messenger RNA (mRNA) was discussed in the last section with a description of the
post-transcriptional modification that convert the pre-mRNA in the nucleus to
mature-mRNA in the cytoplasm.
Transfer-RNA (tRNA) was introduced in the section on the structure of DNA and
RNA and its production and post-transcriptional modification were briefly described in
the previous section. As seen in the illustration below, tRNA is produced as a primary
transcript in a linear primary structure. Following processing the tRNA folds into the
cloverleaf secondary structure. In this structure it is easy to see the 3’ end of the
structure with its ACC sequence that is common to all tRNAs. In addition the bottom loop
in the structure (the anticodon loop) contains the sequence that is complementary and
antiparallel to a sequence on the mRNA. This secondary structure folds to a tertiary
structure in the form of an inverted L. The 3’ end is at one end of the structure and is
the site that will carry a specific amino acid that corresponds to the sequence of
nucleotides in the anticodon at the other end of the molecule. The process by which the
correct amino acid is added to the correct tRNA is referred to as the charging of the
tRNA. The formation of the charged tRNA is a two-step process as depicted in the
"Charging" animation below.
The Following Learn by Doing contains an animation describing the tRNA structures.
-
Ribosomes are quaternary complexes of rRNA and proteins. When they are not
translating RNA to protein, the ribosomes exist as two separated subunits: a large
(50S) subunit and a small (30S) subunit as depicted in the figure below. During the
formation of the 70S initiation complex that starts translation, the two subunits
come together to form the complete ribosome.
 50S and 30S subunits 50S and 30S subunits
|
 70 S Initiation Complex 70 S Initiation Complex
|
-
Initiation Factors are a set of proteins used to facilitate the specific
binding of mRNA to the small subunit of the ribosome.
Protein Synthesis
The following animation describes the formation of the initiation complex, the elongation
of the peptide during synthesis and the termination of the synthesis at the termination
or stop codon. The process is fundamentally the same for prokaryotes and eukaryotes. The
one notable exception is in the binding of the mRNA with the small subunit of the
ribosome during formation of the initiation complex. Prokaryotes do not have a 5’cap to
identify the end of the mRNA but they do have a consensus sequence called the
Shine-Dalgarno-Sequence at the 5’ end of the mRNA. This sequence is used to align the
mRNA with the appropriate site on the ribosomal subunit. In eukaryotic organisms the
5’cap provides the alignment along with multiple initiation factors. Following formation
of the initiation complex, the synthesis of protein follows the same sequence in both
organisms.
The Consequence or Error
During both replication and transcription errors can be made in the incorporation of the
correct bases during the complementary copying of the DNA. In a previous section a
discussion of the methods used by the system to minimize these errors has been
presented. However, even with editing functions, some errors are generated and result in
changing of the reading code used during translation to a protein sequence. These
changes are referred to as mutations and can be categorized into four different
categories described by the end result of the mutation or change.
-
Silent mutation – Because of the degeneracy of the genetic code, multiple
triplet codes represent the same amino acid. Thus, a change in the sequence for a
code may not result in a change in amino acid. This particularly applies to the
wobble or third base in a triplet code sequence. Thermophylic bacteria that operate
optimally at high temperatures have optimized the use of the degeneracy to include
codons with high G and C content to increase the thermal stability (higher melting
temperature) of the resulting duplex DNA.
-
Missense mutation – Some mutations result in the changing of a codon to
represent a different amino acid. Missense mutations that change the amino acid can
have no effect on the structure of a protein or may have a dramatic effect on the
folding of a protein causing loss of function.
-
Nonsense mutation – Another class of mutations may result in the changing
of a codon to one of the three termination or stop codons. This will result in
premature stop in the translation of a protein. Such a mutation at the beginning of
the translation sequence will generally result in total loss of function which some
nonsense mutations near the normal termination of the coding sequence may have
little or no effect on the function of the resulting protein.
-
Frame-shift mutations – Occasionally a mutation will occur that results in
either the insertion or deletion of an extra base in the coding sequence. This type
of mutation results in a shifting of the “reading” frame for the triplet codes after
the insertion or deletion and production of a meaningless sequence of protein after
that point. Again depending on where the insertion or deletion occurs, the effect
can vary on the final product. It should also be noted that this reading frame shift
will also change the position of the termination codon giving either a premature
termination of the translation or an extended coding sequence.
Living organisms consume food, in the form of carbohydrates, fats, and amino acids, to live.
The process of metabolism breaks these complex biomolecules into simple molecules with the
release of energy. The most common form of energy is adenosine triphosphate (ATP). This high
energy phosphorylated compound is used as a energy source in many cellular processes and for
bio-mechanical functions, such as muscle contraction or ion transport across membranes. The
resultant compounds and energy released from food are used to synthesize complex molecules for
the specialized needs of the cell or organism.
The transformation of ingested food to energy and simple compounds, or the synthesis of
complex molecules, is performed by a series of enzymatic reactions. Collectively, these
enzymes, and their substrates and products, are referred to as a metabolic pathway.
Degradative or catabolic pathways generally release energy and electrons from
oxidative processes, i.e.
 Metabolism of Glucose
Metabolism of Glucose
the electrons, which are carried on organic electron carriers, can be used to generate
additional energy, or can be used for synthetic purposes.
Synthetic, or anabolic consume energy and are generally reductive, requiring
electrons.
In this section of the course we will investigate in some detail the production of energy
from glucose and how this process is regulated to maintain homeostasis. Since the entire
process of metabolism is complex, it is useful to discuss the general features of the
metabolic pathways that are involved in converting sugars, amino acids, and fats to energy.
Important features of pathways include:
- input and output compounds,
- the cellular location of the pathway,
- the type of energy the pathway produces.
-
Sugars are degraded as follows:
-
Glycolysis is the first pathway used to degrade sugars and it is located in
the cytoplasm. The simple monosaccharide, glucose, is considered to be the
entry point to this pathway. The three carbon keto acid, pyruvate, is the final
product of glycolysis. Glycolysis generates a small amount of energy in the form of the
high energy compound ATP and high energy electrons, carried on organic electron
carriers. These electrons are brought into the mitochondria.
- The citric acid cycle, or tricarboxylic acid cycle (TCA), is the second
pathway in the degradation of sugars. It is located in the matrix of the mitochondria.
Pyruvate enters the TCA cycle by loss of CO2 to produce a key metabolic
intermediate, acetyl-CoA. The remaining two carbons from pyruvate are lost as
carbon dioxide in the TCA cycle. The energy released by these oxidations is stored on
high-energy electron carriers.
- The electron transport chain consists of a number of proteins that exist as
four distinct complexes in the inner membrane of the mitochondria. This pathway takes
the electrons from the high energy electron carriers and deposits them on oxygen,
generating water. This releases energy, which is used to produce a high concentration of
protons, or a proton gradient across the inner mitochondrial membrane.
-
ATP synthase, which is final step in energy generation, utilizes the proton
gradient to synthesize ATP. This multi-protein enzyme complex also resides in the inner
mitochondrial matrix.
-
Amino Acids are largely degraded by entry into the TCA cycle, followed by
electron transport, and finally ATP synthesis from the proton gradient.
-
Fats, in the form of triglycerides, are first broken down into glycerol and fatty
acids. The fatty acids are oxidized by the beta-oxidation pathway, which coverts the carbon
in the fatty acid to Acetyl-CoA. The acetyl-CoA enters the TCA cycle, this is again followed
by electron transport, and then ATP synthesis.
The location and connections between these degradative pathways is shown below:
OVERVIEW OF METABOLISM. Click the radio buttons to outline in yellow the four key pathways: glycolysis, the TCA cycle, electron
transport, and ATP synthesis and show the flow of carbon atoms, electrons, and protons through each pathway.
Glucose that is brought into the cell
via the glucose transporter can suffer two fates, oxidation or storage as glycogen.
Oxidation occurs in glycolysis and the TCA cycle, releasing the carbon atoms in glucose as
CO2. Note that oxygen is not used until the end of the electron transport
chain. High energy electrons, symbolized as orange balls are carried on organic electron
carriers to the electron transport chain. As these electrons move through the four
complexes, protons are pumped from the mitochondrial matrix across the inner mitochondrial
membrane. As these protons flow back through the membrane via ATP synthase, ATP is
generated.
Many of these pathways, with minor modifications, are reversed for synthetic purposes. For
example, glucose can be synthesized from pyruvate, fatty acids from acetyl-CoA, and amino
acids from intermediates in the TCA cycle.
Degradative, or catabolic, pathways generally produce energy. They usually begin
with a number of different compounds, each of which represents a branch at the beginning of
the pathway. These branches meet at a common intermediate, and the remaining section of the
pathway is usually a linear segment. In this way a number of complex compounds are converted
to a common intermediate, reducing the number of unique steps in the degradation of complex
molecules.
Synthetic, or anabolic, pathways generally consume energy. They usually consist of
an initial linear segment, followed by branching to complex compounds at the end of the
pathway. This strategy allows the use of common simple starting materials for the synthesis of
a number of complex molecules.
Common features of all metabolic pathways are:
- They contain multiple intermediates (e.g. compounds A, B, C, ....), with small molecular
differences between the intermediates.
- Each step, or conversion between intermediates, is catalyzed by an enzyme (e.g.
E1).
- The pathway is regulated to optimize the use of resources.
It is possible to reverse the direction of a metabolic pathways, depending on the needs of
the organisms; a degradative pathway can become a synthetic one. Many of the enzymes that
catalyze reactions in one direction can be easily reversed, and thus function in both
pathways. A small number of steps utilize different enzymes in the forward versus the reverse
direction. These enzymes are regulated in a coordinated fashion such that a pathway operates
in only direction at time.
Pathways can be:
A is the substrate for the first enzyme (E1) in the pathway, B is the
product of the enzyme. The final product of the pathways is compound D. Note that this
pathway can be reversed, using compound D to eventually synthesize compound A.
An example of a branched pathway. The direction of the arrows indicate that this
pathways is an anabolic, or synthetic, pathway where complex biomolecules D and F are
synthesized using the simpler molecule A as starting material. In the reverse direction, the
complex molecules D and F would be converted to A, releasing energy.
In this circular pathway, compound A is transformed to compound B by the enzyme 1. A
series of transformations eventually convert B back to A, restarting the cycle.
It is essential that biochemical pathways are regulated, otherwise the cell would waste
resources. Some general properties associated with the regulation of metabolic pathways are
listed below
- At least one step is regulated, often in multiple ways.
- Essentially irreversible steps, i.e. those with large energy changes, are regulated.
Consequently, once the pathway is turned on, metabolites that enter the pathway are
committed to complete the pathway.
- In synthetic and degradative pathways a common step is performed by different enzymes,
each of which are regulated in a coordinated fashion such that only one pathway is on at a
time.
There are five general methods by which the flux through a step in the pathway can be
regulated. These methods differ in how rapidly they can respond to changes in the environment.
Each of these methods is discussed below, with the more rapid form of regulation at the top of
the list.
-
Substrate availability. For many enzymes, the intracellular level of substrate is
below the level that half saturates the enzyme. Therefore, if the substrate concentration
increases the rate of the reaction will increase due to the formation of additional
enzyme-substrate complexes. If the substrate concentration is very low, the increase in the
rate of the reaction is almost equal to the increase in substrate concentration.
-
Product inhibition. In this case the product of the reaction inhibits the enzyme
that generated it, preventing the accumulation of intermediates in a pathway. The product is
a competitive inhibitor of the enzyme that just created it.
Question: How do product (competitive) inhibitors actually inhibit the reaction of the
substrate with its enzyme? Complete the following Activity and then enter you
answer on the My Response box below it.
Biochem version
new modern bio version version
old version
-
Feedback inhibitors.In this case, a compound that is further down the pathway, or
even a compound in a separate pathway, will inhibit a reaction. Again, this prevents an
accumulation of intermediates in a pathway. The binding of the feedback inhibitor usually
causes a change in the shape of the enzyme; a feedback inhibitor is an example of a
non-competitive inhibitor or an allosteric inhibitor.
- Allosteric Binding
-
(Definition)
Allosteric binding causes conformational changes in an enzyme that can either inhibit or activate the enzyme.
Question: How do allosteric regulators differ from feedback inhibitors? Use the following
activity to explore this difference.
-
Enzyme phosphorylation. The addition of phosphate groups to serine, threonine,
and tyrosine residues on enzymes can cause a change in the shape of the active site of the
enzyme. This change in shape is an allosteric effect. The change may lead to a more active
enzyme (allosteric activator) or a less active enzyme (an allosteric inhibitor).
-
Enzyme levels. Recall that the rate of product formation is proportional to the
amount of enzyme. Enzyme levels can be increased by the conversion of inactive forms of the
enzyme to active forms. This form of regulation occurs with digestive enzymes. These are
made in an inactive form in the pancreas, but activated by cleavage in the small intestine.
The levels of enzyme can also be varied by regulating the synthesis of the enzyme. This
method of regulation is common during development, when certain enzymes are only present
during defined developmental stages.
Oxidation-reduction reactions, or Redox reactions are common in metabolic pathways.
Generally, degradative (catabolic) pathways cause the net oxidation of compounds, releasing
energy. In contrast, synthetic (anabolilc) pathways, are generally reductive pathways.
Oxidations involve the loss of electrons.
Reductions involve the gain of electrons.
Here are two mnemonics to help you remember where the electrons go during redox reactions:
-
LEO GER: "leo [the lion] goes grr". Lose electrons oxidation, gain electron
reduction.
-
OIL RIG: Oxidation involves loss, reduction involves gain.
An example of an oxidation is the conversion of iron from its metallic state, Fe0,
to its rusted form, Fe+3, by the loss of three electrons.
 Oxidation and reduction of iron. Metallic iron, Fe0, becomes oxidized to
Fe+3 (otherwise known as rust) by the removal of 3 electrons.
Oxidation and reduction of iron. Metallic iron, Fe0, becomes oxidized to
Fe+3 (otherwise known as rust) by the removal of 3 electrons.
The above reaction is an incomplete description of a redox reaction because it does not
indicate the fate of the electrons that were obtained from iron. Since free electrons
generally cannot exist, all oxidation reactions must be coupled to a corresponding reduction.
Since the above reaction only describes one-half of the reaction it is referred to as a
half-reaction. The oxidation of iron could be coupled to the reduction of copper
ions, which is described by the following half-reaction:
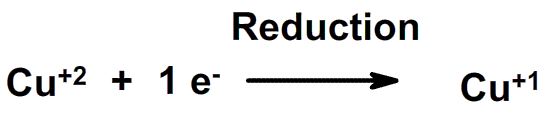 Reduction of copper ion from its +2 state to +1 state by gain of an
electron.
Reduction of copper ion from its +2 state to +1 state by gain of an
electron.
The complete reaction, balanced such that there are no free electrons, is:
 A balanced redox equation. Three copper2+ ions provide a total of three
electrons to oxidize metallic iron to Fe3+. Note that there are no free
electrons.
A balanced redox equation. Three copper2+ ions provide a total of three
electrons to oxidize metallic iron to Fe3+. Note that there are no free
electrons.
The pair of compounds that exchange electrons are often referred to as a redox
couple.
Redox Carriers
In most biochemical redox reactions a total of two electrons are transferred. These
electrons are often transferred as hydrogen atoms, containing a proton and electron. Two
common electron acceptors are NAD+ (nicotinamide adenine dinucleotide) and FAD
(flavin adenine dinucleotide). They both can accept two electrons, giving the reduced
forms NADH and FADH2, respectively. The structure of the oxidized forms of
these compounds are shown below.
 The chemical structures of nicotinamide adenine dinucleotide (NAD+)
and flavin adenine dinucleotide (FAD) are shown. These two compounds are commonly used
as electron acceptors in metabolic pathways. The portion of each molecule that accepts
electrons during the reduction process is highlighted in yellow.
The chemical structures of nicotinamide adenine dinucleotide (NAD+)
and flavin adenine dinucleotide (FAD) are shown. These two compounds are commonly used
as electron acceptors in metabolic pathways. The portion of each molecule that accepts
electrons during the reduction process is highlighted in yellow.
Oxidation of NAD+. In the oxidation of glyceraldehyde to
phosphoglycerate an aldehyde is oxidized to a carboxylic acid and the released electrons
are placed on to NAD+ to form NADH.
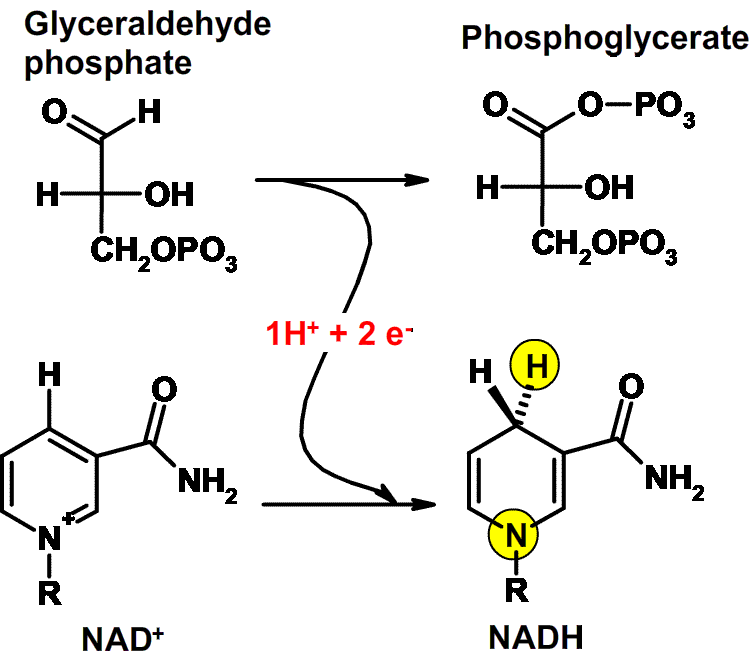 The oxidation of an aldehyde to a carboxylic acid. The two electrons released by
the aldehyde are transferred to NAD+ to make NADH. In this diagram only the
portion of NAD+/NADH that undergoes chemical changes is shown. The remaining
part of the NAD molecule is represented by 'R'.
The oxidation of an aldehyde to a carboxylic acid. The two electrons released by
the aldehyde are transferred to NAD+ to make NADH. In this diagram only the
portion of NAD+/NADH that undergoes chemical changes is shown. The remaining
part of the NAD molecule is represented by 'R'.
Oxidation of FAD. The oxidation of succinate to fumarate, using FAD as an
electron acceptor is another example of a redox reaction found in a metabolic pathway. Two
hydrogen atoms (= two electron plus two protons) are removed from succinate and placed on
FAD, producing fumarate and FADH2, oxidizing a carbon-carbon single bond to a
double bond.
 The oxidation of an alkane to an alkene. The two electrons released by the alkane
are transferred to FAD to make FADH2. In this diagram only the portion of
FAD/FADH2 that undergoes chemical changes is shown. The remaining part of
the FAD molecule is represented by 'R'.
The oxidation of an alkane to an alkene. The two electrons released by the alkane
are transferred to FAD to make FADH2. In this diagram only the portion of
FAD/FADH2 that undergoes chemical changes is shown. The remaining part of
the FAD molecule is represented by 'R'.
Balancing Redox Reactions
It is often difficult to determine from the structure of an organic compound whether it
has been oxidized or reduced in a reaction. For example, the addition of a water molecule
to an double bond (alkene) appears to be a redox reaction because an -OH group has been
added .
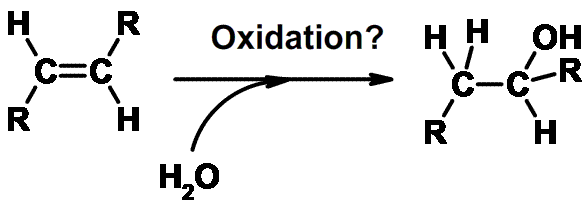 The addition of water to a double bond is a common reaction in many pathways. Is
it a redox reaction?
The addition of water to a double bond is a common reaction in many pathways. Is
it a redox reaction?
The rules for balancing redox reactions are as follows:
- Make the number of oxygen atoms in the reactant and product equal by adding the
appropriate number of water molecules to one side of the reaction or the other.
- Use H+, or H+ + e-, or e- to balance
hydrogen atoms and/or charge.
- A redox reaction has occurred if electrons are consumed or released
The above reaction is balanced as is, and is therefore not a redox reaction.
Try the following mini-tutor to test your skill at assessing whether a redox reaction has
occurred.
The operation of a metabolic pathway produces (catabolic) or consumes (anabolic) energy.
There are a number of different forms of energy storage that are found in metabolic pathways.
These include:
-
Phosphorylated Compounds Nucleoside triphosphates, such as adenosine triphosphate
(ATP) are commonly used to store energy. The addition of an inorganic phosphate group to a
nucleotide diphosphate to form the triphosphate requires approximately 30 kJ/mol of energy.
The reaction showing the synthesis of ATP from ADP and phosphate is pictured below.
 Phosphorylation
of adenosine diphosphate (ADP) produces adenosine triphosphate
(ATP). This reaction requires the input of approximately 30
kJ/mol. Formation of ATP occurs when the negatively charged phosphate
group on ADP attacks the electropositive phosphate in inorganic
phosphate, forming a phosphate anhydride bond with the release of water
(red arrow).
ATP can be used to phosphorylate other nucleoside diphosphates with essentially no
input of energy, for example:
The myth of high-energy phosphate bonds. When a
phosphate is released from ATP to form ADP, about 30
kJ/mol of energy is released. It is often stated, incorrectly,
that the bond that is broken
is "high-energy". In fact, its energy is no different than any
other phosphate bond of the
same type. The release of energy is due to the fact that the
products, ADP and inorganic phosphate, are lower in energy than
ATP by 30 kJ/mol. One reason that ATP is higher in energy is
due to charge repulsion between the negatively charged phosphate groups.
Once the phosphate group is removed, the unfavorable repulsion
disappears.
Phosphorylation
of adenosine diphosphate (ADP) produces adenosine triphosphate
(ATP). This reaction requires the input of approximately 30
kJ/mol. Formation of ATP occurs when the negatively charged phosphate
group on ADP attacks the electropositive phosphate in inorganic
phosphate, forming a phosphate anhydride bond with the release of water
(red arrow).
ATP can be used to phosphorylate other nucleoside diphosphates with essentially no
input of energy, for example:
The myth of high-energy phosphate bonds. When a
phosphate is released from ATP to form ADP, about 30
kJ/mol of energy is released. It is often stated, incorrectly,
that the bond that is broken
is "high-energy". In fact, its energy is no different than any
other phosphate bond of the
same type. The release of energy is due to the fact that the
products, ADP and inorganic phosphate, are lower in energy than
ATP by 30 kJ/mol. One reason that ATP is higher in energy is
due to charge repulsion between the negatively charged phosphate groups.
Once the phosphate group is removed, the unfavorable repulsion
disappears.
-
Reduced redox carriers.The oxidation of metabolytes usually produces energy, If
this energy was not captured in some way, it would be lost as heat. The reduced form of
redox carriers, such as NADH and FADH2, are higher in energy than their
corresponding oxidized forms, capturing the energy that would otherwise be lost as heat. For
example, the oxidation of isocitrate to ketoglutarate releases approximately 70 kJ/mol, 60
of which is captured by converting NAD+ to NADH.
-
High energy thioesters are often produced by oxidative steps. For example, the
energy released by the oxidation of an aldehyde is stored in both the reduced form of
NAD+ as well in a thioester. The hydrolysis of the thioester can be used to
synthesize nucleoside triphosphates or to facilitate the formation of carbon-carbon bonds,
as shown below.
 The oxidation of the aldehyde to the thioester is highlighted in green. CoA is
coenzyme A, a nucleotide containing cofactor that is an essential co-substrate for many
reactions. Part of the energy of this oxidation, 60 kJ/mol, is captured by the formation
of NADH. The energy stored in the thioester can be used to either phosphorylate GDP to
form GTP, capturing another 30 kJ/mol. In the case of acetyl-CoA (lower diagram) the
thioester facilitates the attachment of the acetyl group to oxaloacetate to form
citrate, the first compound in the TCA cycle.
The oxidation of the aldehyde to the thioester is highlighted in green. CoA is
coenzyme A, a nucleotide containing cofactor that is an essential co-substrate for many
reactions. Part of the energy of this oxidation, 60 kJ/mol, is captured by the formation
of NADH. The energy stored in the thioester can be used to either phosphorylate GDP to
form GTP, capturing another 30 kJ/mol. In the case of acetyl-CoA (lower diagram) the
thioester facilitates the attachment of the acetyl group to oxaloacetate to form
citrate, the first compound in the TCA cycle.
-
Proton gradient The transfer of electrons from NADH and FADH2 to
oxygen to form water during the electron transport chain, provides energy for the pumping of
protons across the inner mitochondrial membrane. This is equivalent to pumping water up hill
to fill a reservoir. As the protons flow back through the membrane, the energy released is
used to generate ATP, in much the same way water generates electricity in a hydroelectric
plant.
Free Energy and Spontaneity
Pathways accomplish the net conversion of the starting compounds to the final product of
the pathway. During the normal operation of a pathway there is a constant flux of material
through the pathway in one direction. Note that many pathways are reversible and operate
in the forward or reverse direction, depending on the needs of the organism. The direction
of a pathway depends on the energy difference between the starting compounds and final
product of the pathway. The pathway will be spontaneous in the direction that causes a
decrease in the free energy of the system. Reversing the direction of a pathway
requires changing the relative energies of the reactants and products
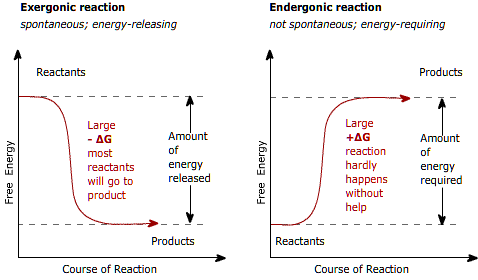
It is useful to have a quantitative way to predict the direction of a pathway given the
current environment in the cell. A method of predicting the direction of a reaction was
devised by J. Willard Gibbs in 1876 and the quantitative parameter that can be used to
predict the direction of a reaction is called the Gibbs free energy. This method
is particularly useful because it can be applied to reactions that are not at equilibrium,
which is the situation encountered during metabolism. Before we can discuss the Gibbs free
energy, we have to discuss standard energies and their relationship to equilibrium
positions of reactions. Keep in mind that these discussions relate to the
thermodynamic properties of pathways, more specifically the relative energy
differences between reactants and products under cellular conditions. The presence of
enzymes simply increases the rate of conversion from reactants to products, the enzyme
cannot alter the relative energies of these compounds.
Equilibria
Consider the simple reaction [A] to [B]. If we start with a system that is pure [A] it
will spontaneously form some [B] until equilibrium is reached. When a system is at
equilibrium, the concentration of products and reactants are constant and it is possible
to write an equilibrium constant for the reaction:
where, KEQ is:
Note that [A] and [B] are at their equilibrium concentrations in this
formula.
Standard Energy
The standard energy change is the energy change when one mole of reactant is converted to
one mole of product, it is the energy difference between reactants and products: Go=Gproductso-Greactantso.
The standard energy change defines the equilibrium position of a reaction through the
following equation:
or
For the simple reaction of A to B, the fraction of the system in state [A] is:
In a similar fashion, the fraction in state [B] is:
You can see from the above equations that if the energy of the products are equal to the
reactants, then the equilibrium concentration of [B] will be equal to [A]. Mathematically,
this can be shown as follows:
If the energy of [B] is equal to [A], then Go = 0. Therefore
KEQ=1, and
fA=1/(1+1) = 0.5 and fB=1/(1+1) = 0.5
Question: What will happen to the relative concentrations of A and B if the energy of form
A is lower than B, what will happen if the energy of form A is higher? Use the
Learn-by-Doing tutorial below to find out.
Gibbs Free Energy
The formula for change in the Gibbs free energy in the reaction for the reaction
direction [A] to [B] is:
Note that in this equation, the concentration of [A] and [B] are not necessarily at their
equilibrium concentrations.
It can be shown that:
- If G less than 0 then the reaction is not at equilibrium and will proceed
spontaneously in the forward direction, A to B.
- If G greater than 0 then the reaction is also not at equilibrium, but will proceed
spontaneously in the reverse direction, B to A.
- If G=0 the reaction is at equilibrium and the concentrations of reactants and
products will not change.
You can explore these relationships in the following tutorial.
In addition to predicting the direction of the reaction, the absolute value of G, ||, is the amount of energy released when the concentrations of reactants and
products change from their non-equilibrium values to their equilibrium values.
Question: What is the Gibbs free energy when the concentration of A
is greater than its equilibrium value? In what direction will the reaction flow? From A
to B or from B to A? Use the Learn-by-Doing tutorial below to find out.
A reaction will be spontaneous in the forward direction if the Gibbs free energy for that
reaction is less than zero. The same reaction, if run in the reverse direction, will have a
positive Gibbs free energy and will therefore be non-spontaneous. If it is necessary to
reverse the direction of this reaction to reverse the direction of the pathway, then the sign
of the GIbbs free energy for the reverse reaction has to become negative. If you recall, the
Gibbs free energy:
consists of two parts, the standard energy change, Go,
and a term that accounts for the non-equilibrium concentrations of [A]
and [B]. Consequently, an unfavorable reaction with a positive
Gibbs free energy can be made
spontaneous by making the sum of these two terms negative by
coupling the unfavorable reaction
to a favorable, energy releasing one. The energy releasing
reaction provides the necessary energy to change the sign of the Gibbs
free energy from positive to negative.
There are two forms of coupling, both may be used to
ensure a negative Gibbs free energy, direct and indirect coupling. Direct coupling reduces the
standard free energy while indirect coupling reduces the second term of the equation. Both of
these methods are described in more detail below.
-
Direct coupling In this case a large negative Go for the reaction is created by directly coupling the unfavorable
reaction to a favorable one on the same enzyme. For example, the phosphorylation of glucose
is very unfavorable if inorganic phosphate is used as the source of phosphate. The standard
energy change of +14 kJ/mol makes the Gibbs energy positive. However, if the unfavorable
reaction is coupled to the conversion of ATP to ADP within the active site of the enzyme
hexose kinase the 30 kJ of energy released from ATP can be utilized to reduce the standard
energy change such that the overall Gibbs free energy becomes negative. The overall reaction
is the transfer of phosphate from ATP to glucose, with an overall standard energy change of
approximately -15 kJ/mol.
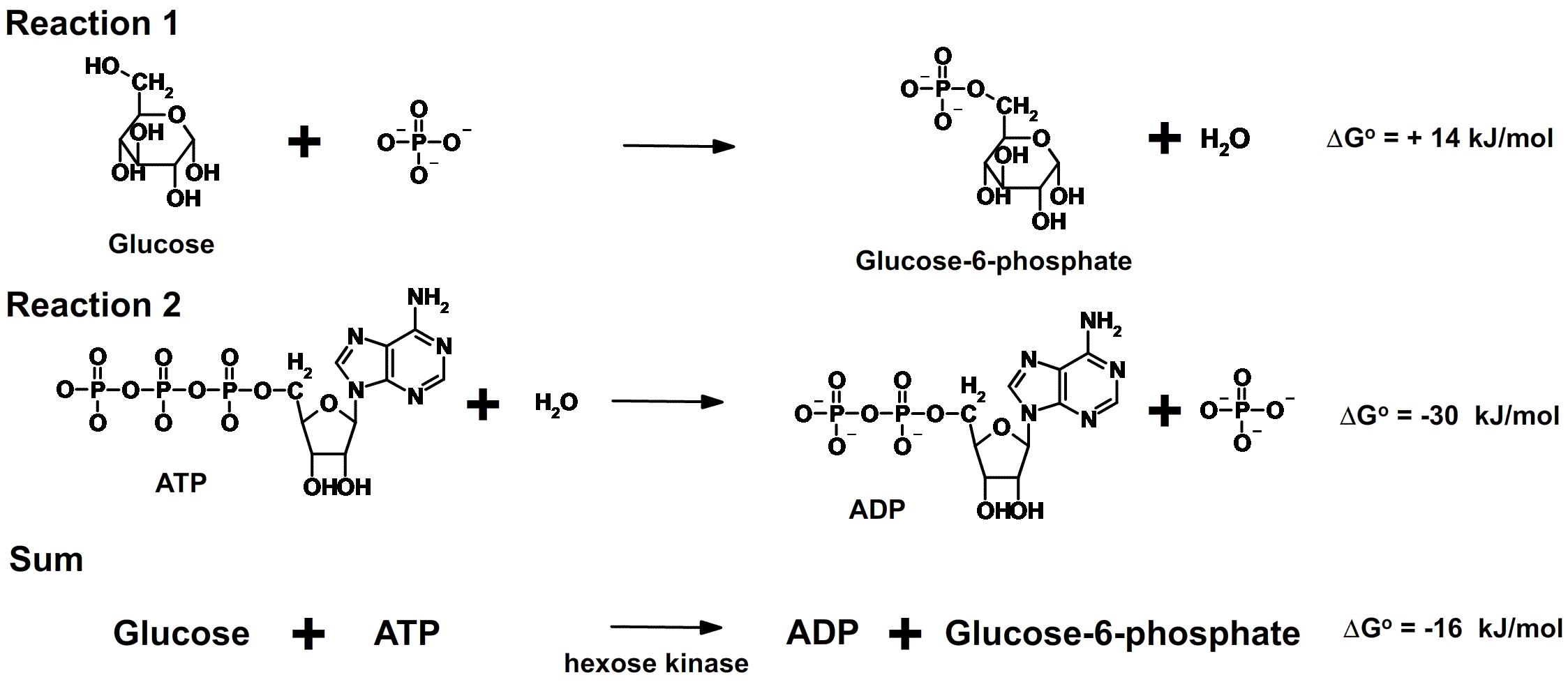 Direct coupling: Reaction 1 and 2 are hypothetical half-reactions that sum to
give the complete reaction at the bottom of the image. In the actual reaction the enzyme
glucose kinase transfers the phospate group directly from ATP to glucose; hydrolysis of
the ATP does not occur. Note that the two half-reactions sum to give the complete
reaction, both in terms of the compounds involved as well as the overall standard energy
change.
Direct coupling: Reaction 1 and 2 are hypothetical half-reactions that sum to
give the complete reaction at the bottom of the image. In the actual reaction the enzyme
glucose kinase transfers the phospate group directly from ATP to glucose; hydrolysis of
the ATP does not occur. Note that the two half-reactions sum to give the complete
reaction, both in terms of the compounds involved as well as the overall standard energy
change.
-
Indirect coupling. The Gibbs free energy can also become negative by either
having a favorable reaction that preceeds the unfavorable one, or a favorable reaction that
follows the unfavorable one. In the first case, the favorable preceeding reaction causes the
concentration of the substrates for the following unfavorable reaction to be higher than
equilibrium, making G negative. For the case of a favorable following reaction, the concentration of
the products of the unfavorable reaction are kept to a level that is smaller than the
equilibrium concentration, again making G negative. This type of coupling between reactions is referred to as indirect
coupling because the coupling between favorable and unfavorable reactions occurs indirectly,
via alteration of concentrations of reactants and products.
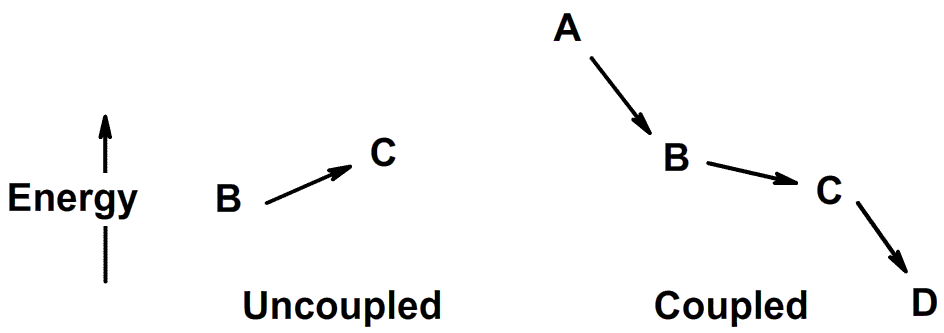 Indirect coupling: In the absence of coupling (left) the conversion of [B] to [C]
is uphill energetically, and therefore not favorable. If the preceeding step ([A] to
[B]) is very favorable, the concentration of [B] increases to a level above its
equilibrium level, which decreases the Gibbs free energy for step [B] to [C].
Alternatively, if the following step ([C] to [D]) is favorable, the concentration of [C]
will be lower than its equilbrium level, which also decreases the Gibbs free energy for
the step [B] to [C].
Indirect coupling: In the absence of coupling (left) the conversion of [B] to [C]
is uphill energetically, and therefore not favorable. If the preceeding step ([A] to
[B]) is very favorable, the concentration of [B] increases to a level above its
equilibrium level, which decreases the Gibbs free energy for step [B] to [C].
Alternatively, if the following step ([C] to [D]) is favorable, the concentration of [C]
will be lower than its equilbrium level, which also decreases the Gibbs free energy for
the step [B] to [C].
There are thousands of different enzymes in any cell. Most enzymes bind a specific substrate,
perform a simple chemical change on that substrate, and then release a product.
Each enzyme has a unique name. Usually, the name of an enzymes is systematic, but many exceptions
exist. Almost all enzyme names have the suffix ase, indicating that they are an enzyme, for
example kinase, lyase, phosphatase, etc. Usually, but unfortunately
not always, the name of the enzyme is derived from the nature of the chemical change that it
catalyzes. For example, an enzyme that oxidizes its substrate is referred to as a
dehydrogenase because it removes hydrogens atoms during the oxidation process.
To further clarify the name of the enzyme, the name of the substrate or product is often included in
the name. For example, the enzyme that oxidizes succinate is called succinate dehydrogenase.
Keep in mind that most reactions in pathways are reversible, so the name may describe the
reverse reaction. Lastly, in cases when enzymes bind more than one substrate, the name can
also suggest the co-substrate. For examples, dehydrogenases will use NAD+ or FAD as
co-substrates.
Important Classes of enzymes
Although there is a large number of enzyme catalyzed reactions in a cell, the following list
describes most of the activities that are found in metabolic pathways.
Kinase. A kinase transfers a phosphate group from ATP to the substrate. Kinases are
used when direct coupling is required to reduce the Gibbs free energy of the reaction.
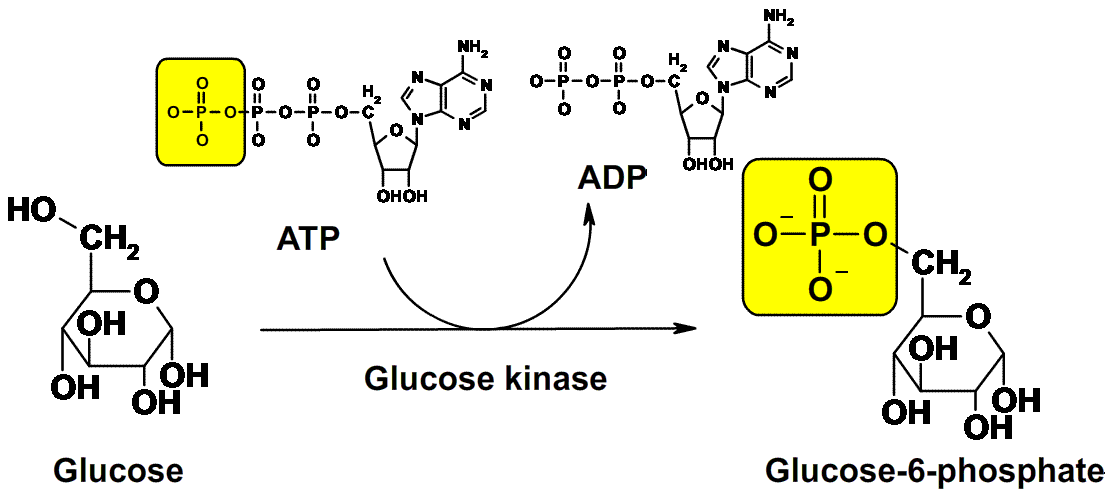 Phosphorylation of glucose by glucose kinase. Note that the source of phosphate is
ATP, not inorganic phosphate.
Phosphorylation of glucose by glucose kinase. Note that the source of phosphate is
ATP, not inorganic phosphate.
Alternatively, a kinase may be involved in regulation of enzymes by transferring a phosphate
from ATP to a Serine, Threonine, or Tyrosine on the enzyme that is being regulated. The
phosphorylated form of the enzyme may be active or inactive.
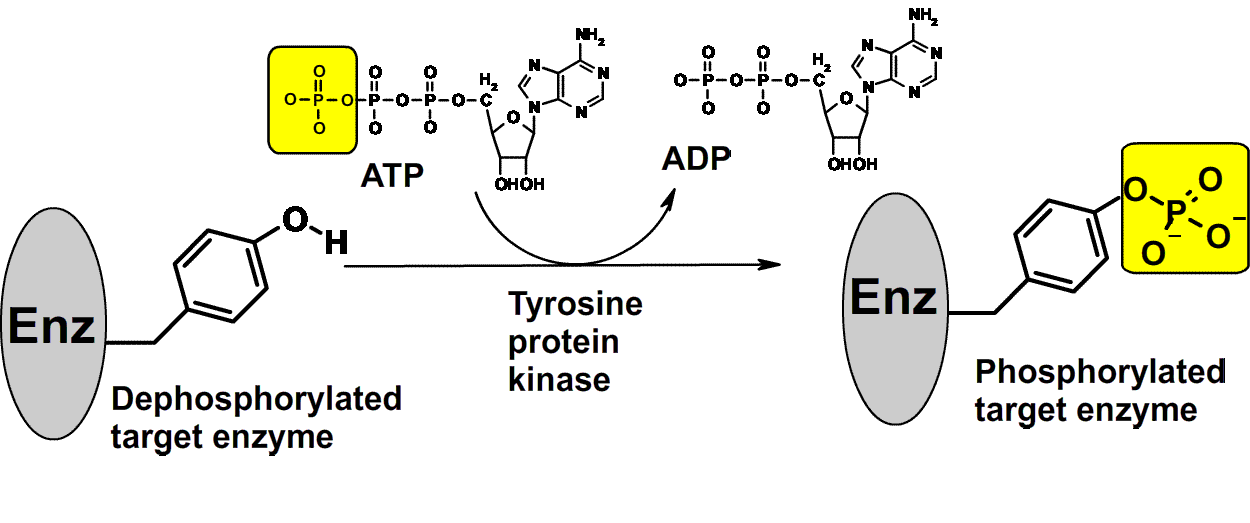 A protein kinase phosphorylates an enzyme, causing it to change its state from active
to inactive or from inactive to active.
A protein kinase phosphorylates an enzyme, causing it to change its state from active
to inactive or from inactive to active.
Phosphatase. A phosphatase removes a phosphate group by a hydrolysis reaction,
producing inorganic phosphate. ADP or ATP are not involved in the reaction.
 A phosphatase uses water to remove a phosphate group from its substrate. Phosphatases
can act on small molecules, as shown above, or on phosphorylated proteins.
A phosphatase uses water to remove a phosphate group from its substrate. Phosphatases
can act on small molecules, as shown above, or on phosphorylated proteins.
Dehydrogenase. As the name suggests, enzymes of this group transfer hydrogen atoms
from the substrate to an electron acceptor, such as NAD+ or FAD . Therefore they
are redox enzymes since removal of hydrogen atoms is the equivalent to removal of electrons.
The name is applied to both oxidation and reduction reactions.
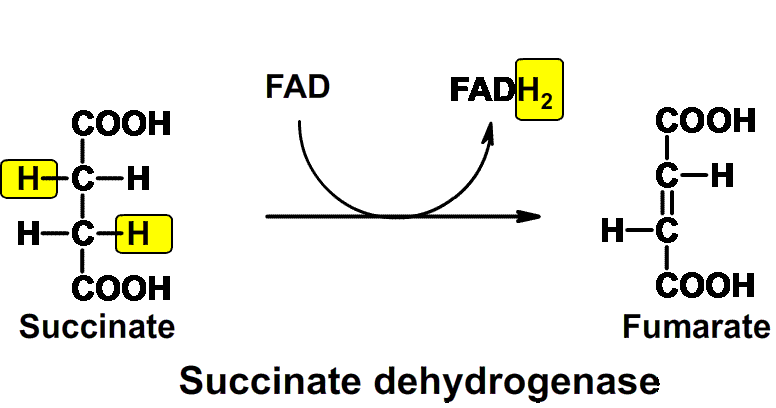 Succinate dehydrogenase, an example from the citric acid (TCA) cycle, is shown. FAD
is the obligatory co-substrate, accepting electrons in this case. Note that this enzyme
could also have been called fumarate dehydrogenase.
Succinate dehydrogenase, an example from the citric acid (TCA) cycle, is shown. FAD
is the obligatory co-substrate, accepting electrons in this case. Note that this enzyme
could also have been called fumarate dehydrogenase.
Isomerase. This class of enzymes rearrange functional groups on their substrates,
releasing a product that has the same number of atoms as the substrate, but is an isomer of
the original substrate. Unfortunately, the descriptive word "isomerase" is often omitted from
their name.
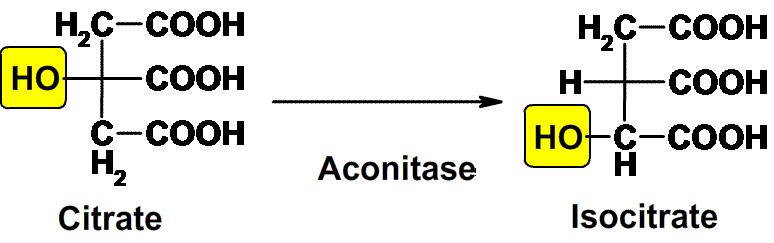 Aconitase is an isomerase that functions in the citric acid cycle to convert citrate
to isocitrate. Note that the composition of both substrate and product are the same, but the
chemical structure is not.
Aconitase is an isomerase that functions in the citric acid cycle to convert citrate
to isocitrate. Note that the composition of both substrate and product are the same, but the
chemical structure is not.
Hydratase. This reaction type, which has a number of idiosyncratic names in
different pathways, adds water to a double bond.
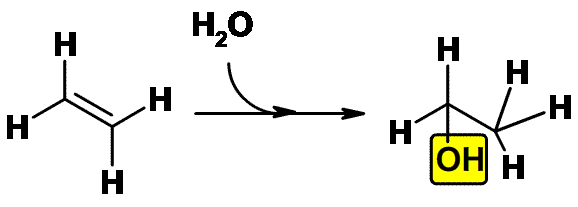 The addition of water to a double bond to produce an alcohol.
The addition of water to a double bond to produce an alcohol.
Synthetase. These enzyme are responsible for the synthesis of more complex molecules
from simpler substrates. For example, ATP synthase generates ATP from ADP and inorganic
phosphate, a process that is driven by a proton gradient across a membrane. Citrate synthase
catalyzes the following reaction, which is the first in the citric acid (TCA) cycle.
 Citrate synthase combines acetyl-CoA and oxaloacetate to form citrate. The source of
energy in this case is the high energy thio-ester in acetyl-CoA.
Citrate synthase combines acetyl-CoA and oxaloacetate to form citrate. The source of
energy in this case is the high energy thio-ester in acetyl-CoA.
Glycolysis is the first pathway that occurs in the metabolism of carbohydrates. This
pathway occurs in the cytosol of all cells, i.e. it is a highly conserved and ubiquitous
pathway. Although many different disaccharides are broken down into monosaccharides and
enter the glycolysis, we will focus on how glucose is metabolized in this section. The
entry points of other sugars will be discussed in the module on integrated
metabolism.
An overall summary of glycolysis is depicted below:
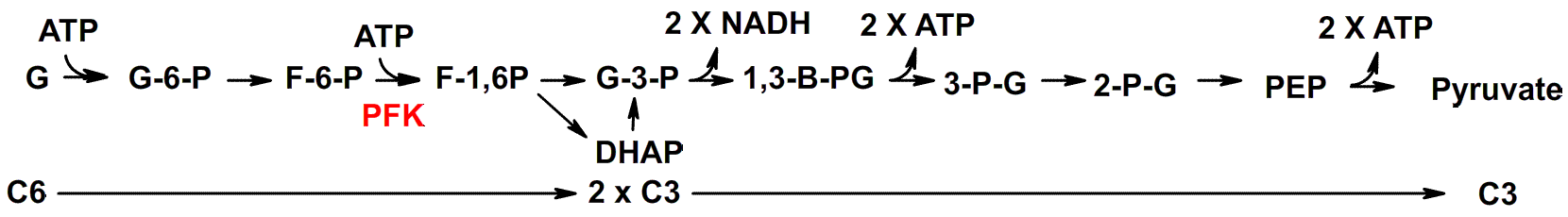 An overview of glycolysis. The intermediates in the pathway are shown near the
top of the figure, along with a summary of energy consumption and production by the
pathway. The molecular structure of these compounds can be seen in the learn-by-doing
exercise below. ADP and NAD+ have been omitted in this diagram for clarity.
The lower part of the figure shows the change in the carbon skeleton in the pathway. The
step following F-1,6-P splits the 6 carbon fructose into two three carbon compounds,
both of which proceed down the pathway to form pyruvate. The key regulatory step, the
enzyme phosphofructose kinase, is indicated as PFK.
An overview of glycolysis. The intermediates in the pathway are shown near the
top of the figure, along with a summary of energy consumption and production by the
pathway. The molecular structure of these compounds can be seen in the learn-by-doing
exercise below. ADP and NAD+ have been omitted in this diagram for clarity.
The lower part of the figure shows the change in the carbon skeleton in the pathway. The
step following F-1,6-P splits the 6 carbon fructose into two three carbon compounds,
both of which proceed down the pathway to form pyruvate. The key regulatory step, the
enzyme phosphofructose kinase, is indicated as PFK.
Key features of Glycolysis:
- Glucose, a six carbon hexose, is the input compound.
- Pyruvate, a three carbon keto-acid is the output. There is no loss of carbon in
glycolysis, so two pyruvates are produced/glucose. Pyruvate is further oxidized in the
TCA cycle.
- Two ATP molecules are produced/glucose. Note that the energy content of two ATP
molecules is required to initiate the pathway. Four ATP (2/pyruvate) are produced later
in the pathway. Consequently, the net yield is two ATP.
- Two NADH molecules are produced. A single oxidation step produces one molecule of
NADH/pyruvate. The energy stored in NADH is extracted during electron transport.
- The key regulatory step is the addition of the second phosphate to fructose, by the
enzyme phosphofructose kinase
Capture of Glucose by the Cell
Glucose from the outside of the cell is transported across the cell membrane by a
multi-subunit protein called the glucose transporter. This enzyme catalyzes the diffusion
of glucose across the membrane without the input of energy. Consequently, spontaneous flow
of glucose into the cell can only occur if the concentration of glucose is lower in the
cell than outside the cell. The Gibbs free energy for the transport of glucose is:
Although this expression for the Gibbs free energy appears different than previous
expressions, it is really the same equation. The "product" of the reaction is glucose
inside the cell and the "reactant" is glucose outside the cell. The difference in standard
free energy, Go, is zero because the reactants and products are the same compound,
differing only in their location. If the concentration of glucose outside the cell is
higher than the concentration inside the cell, then the Gibbs free energy is negative (the
ln of a number less than one is negative) and glucose will spontaneously enter the cell.
If the glucose concentration in both compartments are the same, then =0 and there is no net movement of glucose across the membrane. If the
concentration of glucose inside the cell exceeds the concentration outside then >0 and the reverse reaction, the net movement of glucose out of the cell, will be
spontaneous.
The levels of glucose in the blood vary considerably at times. We will discuss how
glucose levels are regulated in the section on integrated metabolism. The focus here is to
understand how the entry of glucose into the cell is spontaneous, even when the levels of
glucose in the blood can drop to low values. How is it possible to maintain a negative
Gibbs free energy for glucose transport into the cell, even if there are low levels of
glucose outside the cell?
Question: Use the tutorial below to discover how glucose spontaneously enters the cell,
even if the intracelluar concentration exceeds the concentration outside the
cell.
Glycolysis
The following learn-by-doing activity allows you to investigate the chemical and
energetic changes that occur in glycolysis. Once you open up the page, explore it using
the embedded questions to prepare you for the self-assessment at the end of this module.
Question: As you move through the glycolysis pathway, determine which steps become
spontaneous by direct coupling, which by indirect coupling, and how energy is captured
from steps that release large amounts of energy.
Anaerobic Glycolysis
Under anaerobic (oxygen limited) conditions, which occurs in the muscle tissue during
vigorous activity, the NADH produced in glycolysis cannot be reoxidized to NAD+
by electron transport because there is insufficient oxygen to accept electrons. Under
these conditions, the cell runs out of NAD+ and glycolysis will halt and the
cell can no longer produce ATP.
The levels of NAD+ can be restored by using pyruvate as the electron acceptor.
In mammals, lactate is the product of this reaction. In yeast, it is alcohol and the
process of anaerobic glycolysis is referred to as fermentation. The reduction of
pyruvate will oxidize NADH to NAD+, allowing glycolysis to resume.
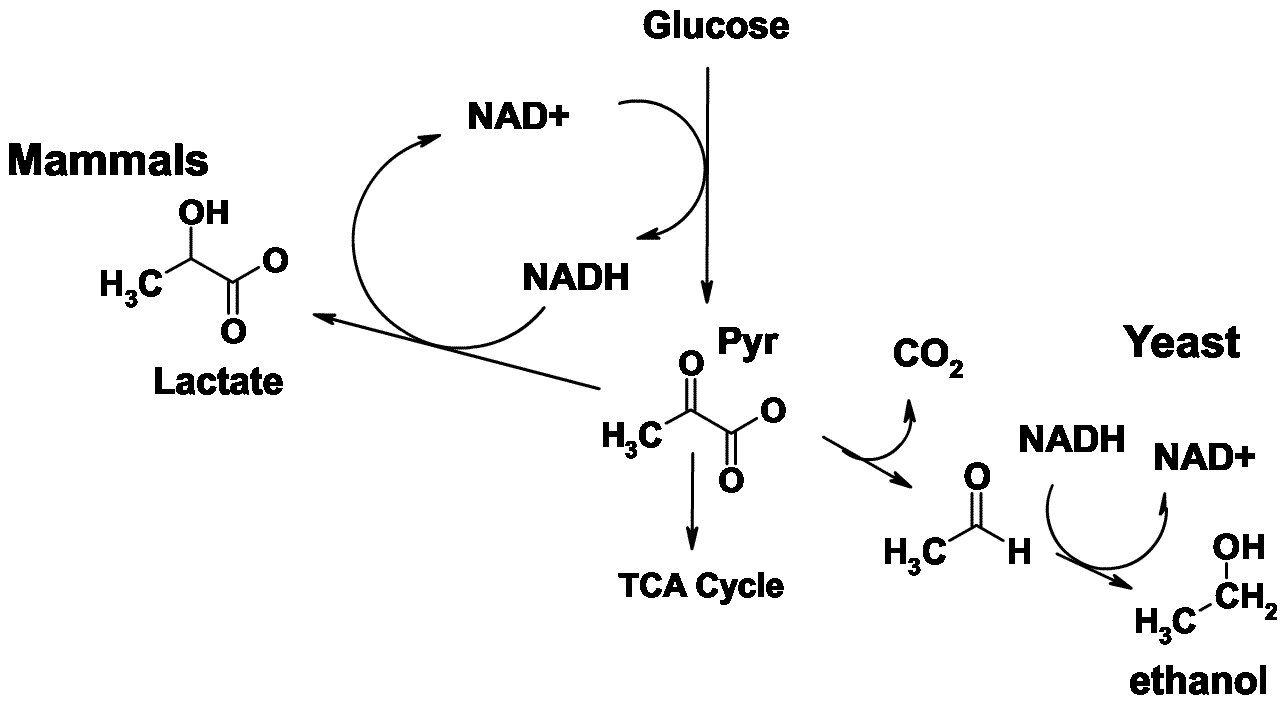 Anaerobic metabolism. Pyruvate can serve as the electron acceptor for NADH. The
reduction of pyruvate to lactate in mammals, or ethanol in yeast, regenerates the
NAD+ required for glycolysis to operate.
Anaerobic metabolism. Pyruvate can serve as the electron acceptor for NADH. The
reduction of pyruvate to lactate in mammals, or ethanol in yeast, regenerates the
NAD+ required for glycolysis to operate.
The lactate that is produced by active muscle tissue is transported to the liver. When
oxygen becomes available, the lactate is reoxidized to pyruvate and it can then be used
for the synthesis of glucose.
Catabolic Process of the TCA Cycle
The tricarboxylic acid (TCA) cycle, also known as the Krebs cycle or the citric acid
cycle, is a central pathway in the metabolism of all organisms. Not only is this pathway
the next step for the oxidation of glucose and fatty acids, it also plays a key role in
amino acid metabolism. All carbons that enter the TCA cycle, regardless of their source,
are fully oxidized to carbon dioxide. The electrons that are released from these
oxidations are stored on NADH and FADH2 for subsequent processing by the
electron transport chain.
Entry of carbohydrates into the TCA cycle occurs by the transport of pyruvate from the
cytoplasm into the mitochondrial matrix. There, the pyruvate undergoes a process called
oxidative decarboxylation to produce a key intermediate in metabolism,
acetyl-CoA, which enters the TCA cycle.
Entry of fatty acids into the TCA occurs directly into acetyl-CoA. During the
process of fatty acid oxidation (beta-oxidation) which also occurs in the mitochondrial
matrix, two-carbon fragments are progressively released from fatty acids and incorporated
into acetyl-CoA.
 Entry of carbohydrates and fats into the TCA cycle. Oxidative decarboxylation of
pyruvate produces acetyl-CoA, releasing one CO2 and NADH. Fatty acid
oxidation yields acetyl-CoA directly. Acetyl-CoA is a high-energy thioester that enters
the citric acid cycle.
Entry of carbohydrates and fats into the TCA cycle. Oxidative decarboxylation of
pyruvate produces acetyl-CoA, releasing one CO2 and NADH. Fatty acid
oxidation yields acetyl-CoA directly. Acetyl-CoA is a high-energy thioester that enters
the citric acid cycle.
The first step of the TCA cycle utilizes the high-energy thioester in acetyl-CoA
to drive the addition of an acetate group to oxaloacetic acid to produce citrate.
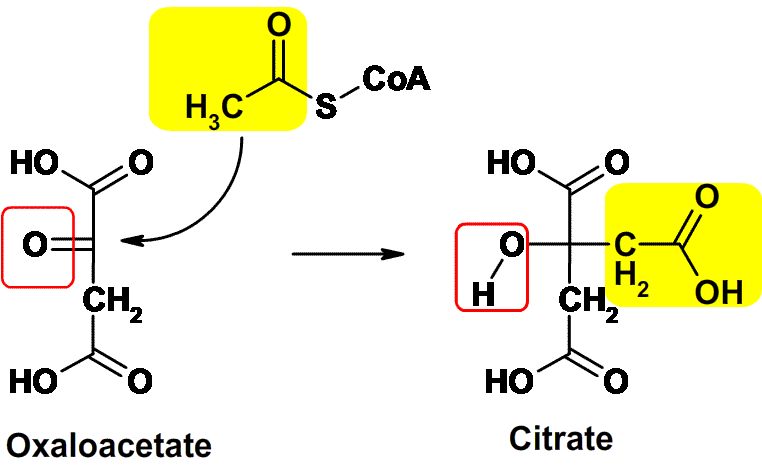 In the first step of the TCA cycle acetyl-CoA donates an acetate group to
oxaloacetate, forming citrate. Note that the ketone group on oxalacetate has become an
alcohol in citrate.
In the first step of the TCA cycle acetyl-CoA donates an acetate group to
oxaloacetate, forming citrate. Note that the ketone group on oxalacetate has become an
alcohol in citrate.
The remaining steps in the TCA cycle convert citrate back to oxaloacetate. This process
produces:
- 2 CO2, from oxidative decarboxylations.
- 3 NADH, all from oxidation of alcohols to ketones.
- 1 FADH2, from the oxidation of an alkane to an alkene
- 1 GTP, from the hydrolysis of a thio-ester.
Given the above information, your task is to deduce the series of reactions that convert
citrate back to oxaloacetate using the following learn-by-doing activity.
Anabolic Role of the TCA Cycle.
Intermediates of the TCA cycle are utilized in the following synthetic
pathways:
- Fatty acids and steroids from acetyl-CoA.
- Amino acids aspartate, asparagine, lysine, isoleucine, methionine from oxaloacetate.
- Amino acids glutamate, glutamine, proline, arginine from alpha-ketoglutaric acid
- Porphyrin, a precursor to heme, from Succinyl-CoA
What major metabolite is missing from this list? Carbohydrates! It
is not possible for mammals
to use carbon atoms derived from the TCA cycle to synthesize
glucose. How glucose is synthesized from pyruvate will be discussed in
the module on integrated metabolism.
If compounds in the TCA cycle are used to synthesize other compounds, such as amino acids, the TCA cycle will
eventually halt due to depletion of oxaloacetate. Consequently it is necessary to "fill
up" the TCA cycle with an "anaplerotic" reaction. In this case, pyruvate is
converted to oxaloacetate by the enzyme pyruvate carboxylase:
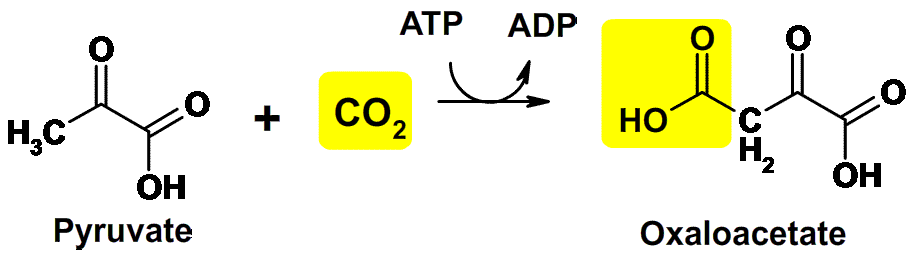 Anaplerotic reaction that generates oxaloacetate from pyruvate. This reaction
replenishes the carbons in the TCA cycle, allowing it to continue to operate under
anabolic conditions.
Anaplerotic reaction that generates oxaloacetate from pyruvate. This reaction
replenishes the carbons in the TCA cycle, allowing it to continue to operate under
anabolic conditions.
The production of ATP from "high-energy" electrons on NADH and FADH2 occurs in two
steps, electron transport followed by ATP synthesis. The energy obtained from electron
transport is stored as a pH gradient across the inner mitochondrial membrane. The combination
of a concentration difference of hydrogen ions and a voltage difference accross this membrane
is often referred to as an electrochemical gradient. The electrochemical gradient is
a form of potential energy that is utilized by the enzyme ATP synthase to convert ADP to ATP.
The coupling between the proton gradient and the chemical synthesis of ATP, was originally
proposed by Dr. Peter Michell in 1961 as the chemiosmotic hypothesis. His theory
proved to be true, leading to a Nobel prize for his work in 1978.
Electron Transport
The electron transport chain consists of four multi-protein complexes that are
contained within the inner mitochondrial membrane. These complexes remove
electrons from NADH or FADH2 molecules that were generated by oxidative
processes in glycolysis and the TCA cycle. These high energy compounds ultimately deposit
their electrons on oxygen, forming water. The energy that is released by electron
transport is stored as a proton gradient across the inner mitochondrial membrane. This
proton gradient is used to drive the synthesis of ATP.
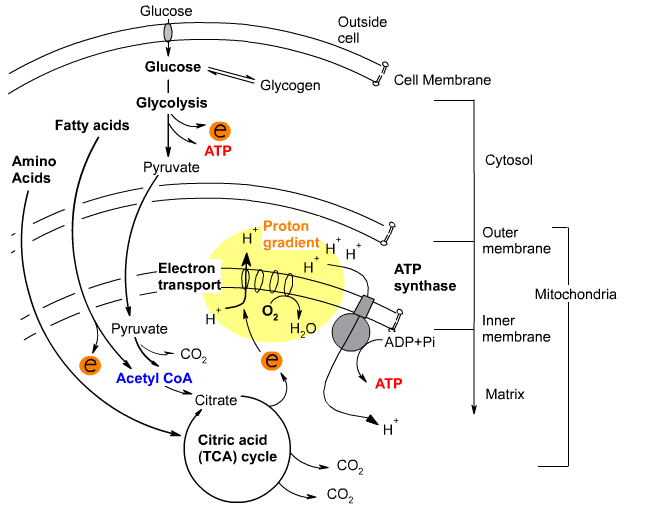 The hilighted region focusing in on the Electron Transport Chain, part
of the complex process of metabolism.
The hilighted region focusing in on the Electron Transport Chain, part
of the complex process of metabolism.
Various compounds are used by the electron transport chain to carry electrons. These
components include:
-
FAD/FADH2, is a cofactor that is tightly bound to the enzymes in
the electron transport complexes.
-
Iron-sulfur centers in the electron transport complexes shuttle electrons by
alternating between Fe+2 and Fe+3.
-
Coenzyme Q is an organic non-polar electron carrier that is dissolved within
the membrane lipids of the inner mitochondrial membrane. It carries electrons between
complex I and III and complex II and III.
-
Cytochrome C is a heme containing protein that is very similar in structure
to myoglobin. It is a water soluble protein found in the inter-membrane space.
Cytochrome C carries one electron at a time from complex III to complex IV.
The organization of the four complexes and electron carriers in the electron transport
chain are illustrated in the following diagram:
The electron transport chain is contained within the inner membrane of the
mitochondria. In addition to the four protein complexes, the location of electron
carriers coenzymeQ and cytochrome C are also indicated. Complex I accepts electrons from
NADH and complex II accepts electrons from succinate. Complex IV is responsible for
transferring the electrons to the final electron acceptor, oxygen. Selecting the button
labeled NADH will show the flow of electrons from NADH to complex IV. The button labeled
FADH2 will show the path of electrons from succinate to FADH2 in
complex II (succinate dehydrogenase), followed by movement of these electrons to complex
IV.
NADH is oxidized by the electron transport chain by the transfer of the two
electrons from NADH to complex I. These electrons are then transferred to coenzyme Q,
followed by transfer to complex III. The electrons are then carried by cytochrome C to
complex IV, where the electrons are used to reduce oxygen to water. The energy that is
released by the transfer of electrons from NADH to water is used to transport a total of
10 protons across the inner membrane. For every pair of electrons, four protons are
transported by complex I, four by complex II and two by complex IV.
The electrons from FADH2 are first processed by complex II. Complex
II is actually succinate dehydrogenase from the TCA cycle. The electron acceptor for
succinate, FAD, is tightly bound to the enzyme. Consequently, it is more correct to
consider that complex II processes the electrons from succinate, first passing them to FAD
and then via iron-sulfur centers to coenzyme Q. The two electrons then follow the same
path as those from NADH2, from coenzyme Q to complex III, and then to complex
IV via cytochrome C, finally to oxygen to produce water. Since electron transport through
complex II does not result in the transport of protons only a total of 6 protons/2
electrons are transported across the inner membrane when succinate is the initial electron
donor.
In summary:
- Electrons from NADH and FADH2 are transferred to oxygen, generating water.
- Electron transport is spontaneous and releases energy. This energy is stored by
transporting protons across the inner membrane.
- Oxidation of NADH results in 10 protons transported across the inner membrane.
- Oxidation of FADH2 results in 6 protons transported across the inner
membrane.
 This link will take you to a
movie providing an alternate representation of the electron transport process.
This link will take you to a
movie providing an alternate representation of the electron transport process.
ATP Synthesis
The energy that has been stored in the proton gradient across the inner mitochondrial
membrane cannot be easily utilized by other processes in the cell. It must be converted to
a more usable source, such as ATP. The enzyme ATP synthase is responsible for converting
the energy stored in the proton gradient to ATP. This enzyme is found in the inner
mitochondrial membrane and it projects into the matrix.
The mechanism of ATP synthase is such that the enzyme generates one ATP molecule every
time 3 protons are transferred back to the matrix from the inter-membrane space. Since the
standard energy for the formation of ATP from ADP and Pi is +30 kJ/mol the transport of
three protons must release at least 30kJ/mol in order to provide enough energy to
synthesize ATP. The amount of energy that is stored in the protein gradient can be
obtained by calculating the Gibbs free energy. If the products are defined as the protons
that have been transported to the matrix, the formula is:
where Z is the change on the transported particle, F is Faraday's constant (96,494
Coulomb/mol) and is the voltage difference across the membrane:
The Gibbs free energy consists of two terms. The first indicates the energy change due to
the difference of proton concentration across the membrane. Its form is analogous to the
formula give for the transport of glucose across the membrane in glycolysis. The second
term accounts for the fact that the energy of a charged particle depends on the voltage.
If there is a voltage difference across the membrane then the energy of the proton will
depend on its location.
Typical proton concentration differences across the inner membrane are approximately 10
fold, with the outside being more acidic. In addition to the concentration difference,
there is also an approximately 100 mV voltage difference across the membrane, with the
inside being more negative. Therefore, the Gibbs free energy change under these conditions
at 300 K is:
Therefore the transport of 3 protons will release 45.9 kJ/mol, which is more than
sufficient to generate a single ATP.
Mechanism of ATP Synthase
The synthesis of ATP utilizing the proton gradient as a source of energy is an example
of direct coupling. The energy released as the protons flow through the enzyme cause a
conformational change in the protein that causes the formation of ATP from bound ADP and
inorganic phosphate.
The enzyme consists of two separable subunits. The Fo subunit is within the
membrane and is responsible for the transfer of protons through the membrane. The
F1 subunit extends into the mitochondrial matrix and is responsible for the
synthesis of ATP. The F1 subunit is composed of three subunits and three subunits. These six subunits from a spherical structure where the and subunits alternate. The subunit extends from the Fo domain through the center of the sphere. The subunit rotates 120 degrees every time three protons flows through the
Fo domain. The sphere is prevented from rotating along with the subunit by the b-subunit, which anchors the sphere to the membrane.
The conformation of the subunits is affected by the relative position of the subunit. Since the gamma subunit rotates 120o, there are three
possible conformations of the subunits:
- A conformation that has low affinity of nucleotides, i.e. neither ATP or ADP bind.
- A conformation that has high affinity for ADP plus inorganic phosphate.
- A conformation that has high affinity for ATP, i.e. ATP is more stable in this
conformation than ADP.
The cycle of ATP synthesis is as follows:
- ADP and inorganic phosphate bind to the subunits that have high affinity for these
compounds.
- A three protons flow through the Fo domain, causing a 120o
rotation of the -subunit.
- The rotation of the -subunit causes a conformational change in the subunits.
- ATP is more stable in the new conformation of the subunits, consequently the bound ADP and inorganic phosphate is spontaneously
converted to ATP.
- Another three protons flow through the Fo domain, causing another
120o rotation of the subunit.
- This rotation causes an additional conformational change of the subunits, generating the conformation that has low affinity for ATP and ADP,
thus the newly synthesized ATP is released.
- A three additional protons flows through the Fo domain, causing another
rotation of subunits. This restores the system to the starting conditions.
Complete rotation of the subunit requires the transfer of 9 protons across the membrane. This generates a
total of 3 ATP molecules since there are three ADP/ATP binding sites on the F1
domain, one associated with each subunit. Consequently, only three protons are required to synthesize one ATP.
Click the green arrow or PLAY button to play the animation.
The storage and metabolism of glucose is controlled at the organ level by hormones. The
hormones glucagon and insulin are secreted by the pancreas during periods of
low or high blood sugar, respectively. Glucagon causes the liver to produce glucose from the
storage polysaccharide glycogen or to synthesize glucose from pyruvate using the
pathway gluconeogenesis. The released glucose enters into the blood and travels to
muscle for oxidation by glycolysis leading to energy production. In contrast, insulin
instructs the liver cell to store the excess glucose in the blood.
The hormone epinephrine,which is produced by the central nervous system in response
to dangerous situations, evokes the same response as glucagon, the release of glucose from the
liver.
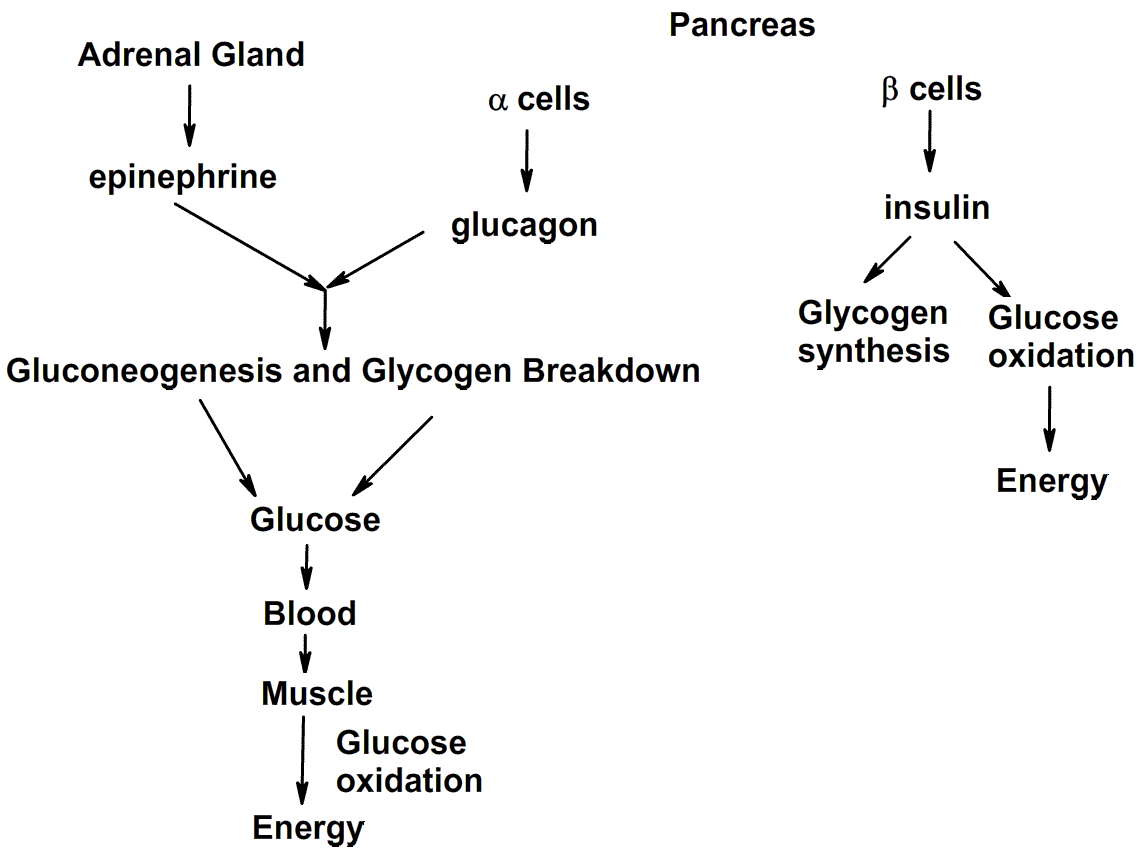 Hormonal control of glucose metabolism. Glucagon and insulin are pancreatic hormones
that regulate blood sugar levels. Epinephrine (adrenaline) is produced by the adrenal gland
in the central nervous system in response to dangerous situations. Glucagon and epinephrine
instruct the liver cell to produce glucose by release from glycogen or by synthesis from
pyruvate via the synthetic pathway gluconeogenesis. The glucose enters the blood and is
oxidized in muscle cells to produce energy. Insulin is secreted when the blood glucose level
is high, instructing the liver to store glucose in glycogen or, if necessary, to oxidize it
to produce energy.
Hormonal control of glucose metabolism. Glucagon and insulin are pancreatic hormones
that regulate blood sugar levels. Epinephrine (adrenaline) is produced by the adrenal gland
in the central nervous system in response to dangerous situations. Glucagon and epinephrine
instruct the liver cell to produce glucose by release from glycogen or by synthesis from
pyruvate via the synthetic pathway gluconeogenesis. The glucose enters the blood and is
oxidized in muscle cells to produce energy. Insulin is secreted when the blood glucose level
is high, instructing the liver to store glucose in glycogen or, if necessary, to oxidize it
to produce energy.
These hormones do not directly affect the enzymes involved in glucose metabolism and storage.
Rather they bind to membrane receptors on the surface of the cell and evoke a conformational,
or allosteric, change in the receptor, transmitting the signal to the inside of the cell.
- The signalling pathway that is activated by the hormones glucagon and
epinephrine begins with the activation of a G-protein by exchange of bound GDP
for GTP. The G-protein then activates adenyl cyclase, an enzyme that converts ATP to cAMP
(cyclic AMP). The cAMP then activates a protein kinase, leading to the
phosphorylation of a number of enzymes in the cell.
- The binding of Insulin to its receptor also causes a conformational change in the
receptor. Subsequent signalling steps do not involve G-proteins and they ultimately lead to
the activation of a protein phosphatase, causing the dephosphoryation of enzymes in
the cell.
Transmitting the Signal to the Inside of the Cell
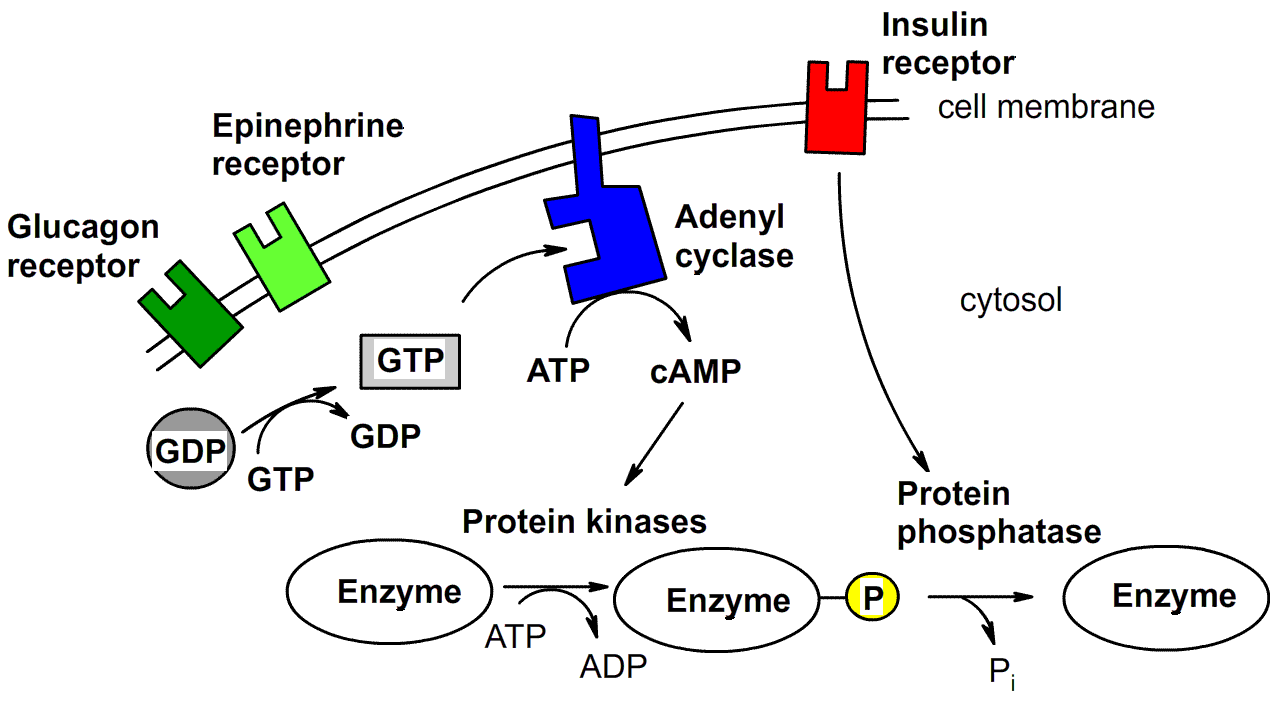 The hormones glucagon, epinephrine, and insulin all bind reversible to receptors on
the cell surface. The binding of glucagon and epinephrine ultimately lead to protein
phosphorylation by the activation of protein kinases while the binding of insulin ultimately
leads to activation of protein phosphatases which remove the phosphate groups from
enzymes.
The hormones glucagon, epinephrine, and insulin all bind reversible to receptors on
the cell surface. The binding of glucagon and epinephrine ultimately lead to protein
phosphorylation by the activation of protein kinases while the binding of insulin ultimately
leads to activation of protein phosphatases which remove the phosphate groups from
enzymes.
Click the green arrow to play the animation.
The phosphorylation of enzymes during periods of glucose demand or the dephosphorylation of
enzymes when there is surplus glucose results in the correct biological response of the liver
cell. Phosphorylation results in the activation of enzymes that cause the release of glucose
from glycogen and also results in the synthesis of new glucose by the anabolic pathway
gluconeogenesis. Dephosphorylation of enzymes causes the activation of enzymes involved in the
storage of glucose in glycogen and can also result in the activation of glucose oxidation for
ATP production.
Since the protein kinases and phosphatases that are involved in the activation of enzymes are
enzymes themselves, amplification of the initial hormone-receptor signal occurs. This is
especially true in the case of the hormones glucagon and epinephrine where additional
amplification occurs due to the production of a large number of cAMP molecules per signalling
event.
Glycogen is a polysaccharide that is used to store glucose in both liver and muscle
cells. In plants, amylopectin serves the same role. Glycogen consists of glucose residues
linked together in a linear chain with -(1-4) linkages. Multiple chains are joined together by the formation of -(1-6) linkages, giving rise to a highly branched polymer. The principle reason for
the storage of glucose as glycogen is to reduce the osmotic effects that would otherwise occur
from the high concentration of glucose in cells.
 Glycogen. The overall structure of a small glycogen molecule is shown on the left. An
enlarged portion of the molecule is shown on the right.
Glycogen. The overall structure of a small glycogen molecule is shown on the left. An
enlarged portion of the molecule is shown on the right.
When needed, glucose is released from glycogen by a reaction catalyzed by the enzyme
glycogen phosphorylase. This enzyme breaks an -(1-4) linkage at the end of glycogen polymer by the addition of an inorganic
phosphate. The product of this reaction, glucose-1-P, can enter glycolysis after conversion to
glucose-6-P. Note that this entry point for glucose skips the first step in glycolysis,
avoiding the need to activate glucose using ATP.
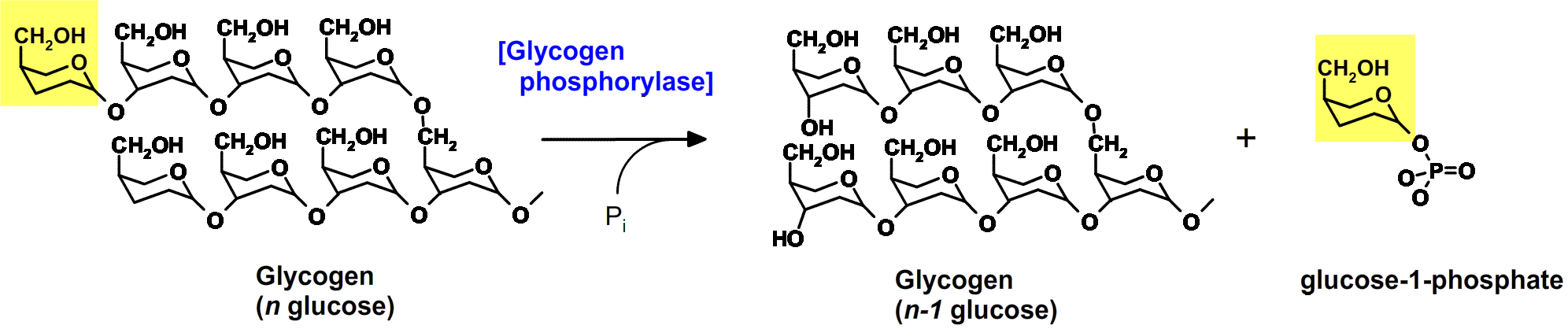 Glycogen degradation occurs by the cleavage of an -(1-4) linkage with inorganic phosphate, catalyzed by the enzyme glycogen
phosphorylase. This is reaction occurs spontaneously, no external energy is required.
Glycogen degradation occurs by the cleavage of an -(1-4) linkage with inorganic phosphate, catalyzed by the enzyme glycogen
phosphorylase. This is reaction occurs spontaneously, no external energy is required.
For storage, glucose is added to the free 4-hydroxyl groups of the glycogen molecule,
elongating the linear chains. Since the release of glucose by phosphorolysis is spontaneous,
the direct addition of glucose to glycogen must be energetically unfavorable. Consequently,
the synthetic reaction requires coupling to an energy releasing reaction to become
spontaneous. The addition of glucose to glycogen proceeds in two steps. First,
glucose-1-phosphate reacts with UTP to produces a high energy intermediate, UDP-glucose, which
in turn is used by the enzyme glycogen synthase to add glucose to glycogen.
 Glycogen synthesis. The incorporation of glucose into glycogen requires two steps. In
the first (I) activation of glucose occurs using the energy from UTP to form UDP-glucose.
Remarkably, the formation of UDP-glucose occurs with a free energy change of nearly zero.
The overall Gibbs free energy of the reaction becomes negative due to the energy released
when pyrophosphate is hydrolyzed to inorganic phosphate. In step II, the high energy
compound UDP-glucose is used to spontaneously add glucose to glycogen. The net energy cost
of adding a glucose to glycogen is two ATPs, one to form glucose-1-phosphate and one to form
UTP from UDP. Since glucose is released from glycogen in a phosphorylated state,
glucose-1-phosphate, the net cost to store glucose in glycogen is one ATP/glucose.
Glycogen synthesis. The incorporation of glucose into glycogen requires two steps. In
the first (I) activation of glucose occurs using the energy from UTP to form UDP-glucose.
Remarkably, the formation of UDP-glucose occurs with a free energy change of nearly zero.
The overall Gibbs free energy of the reaction becomes negative due to the energy released
when pyrophosphate is hydrolyzed to inorganic phosphate. In step II, the high energy
compound UDP-glucose is used to spontaneously add glucose to glycogen. The net energy cost
of adding a glucose to glycogen is two ATPs, one to form glucose-1-phosphate and one to form
UTP from UDP. Since glucose is released from glycogen in a phosphorylated state,
glucose-1-phosphate, the net cost to store glucose in glycogen is one ATP/glucose.
Regulation of Glycogen Synthesis and Degradation: Glycogen synthesis and degradation
is controlled by the phosophorylation state of the enzymes. The phosphorylation levels of
proteins within the cell are directly controlled by response to hormones, as discussed above.
- When there is demand from glucose protein phosphorylation occurs, signalling the release
of glucose from glycogen in the liver. Therefore the phosphorylated state of glycogen
phosphorylase must be the active form of the enzyme so that glucose is released from
glycogen. To insure that both synthetic and degradative pathways are not active at the same
time we would expect that the phosphorylated state of glycogen synthase must be inactive.
- During periods of high blood sugar levels, hormonal signalling causes the storage of
glucose in glycogen by leading to dephosphorylation of enzymes in the cell. This
dephosphorylation activates glycogen synthase and inactivates glycogen phosphorylase, again
insuring that only one direction is active at a time.
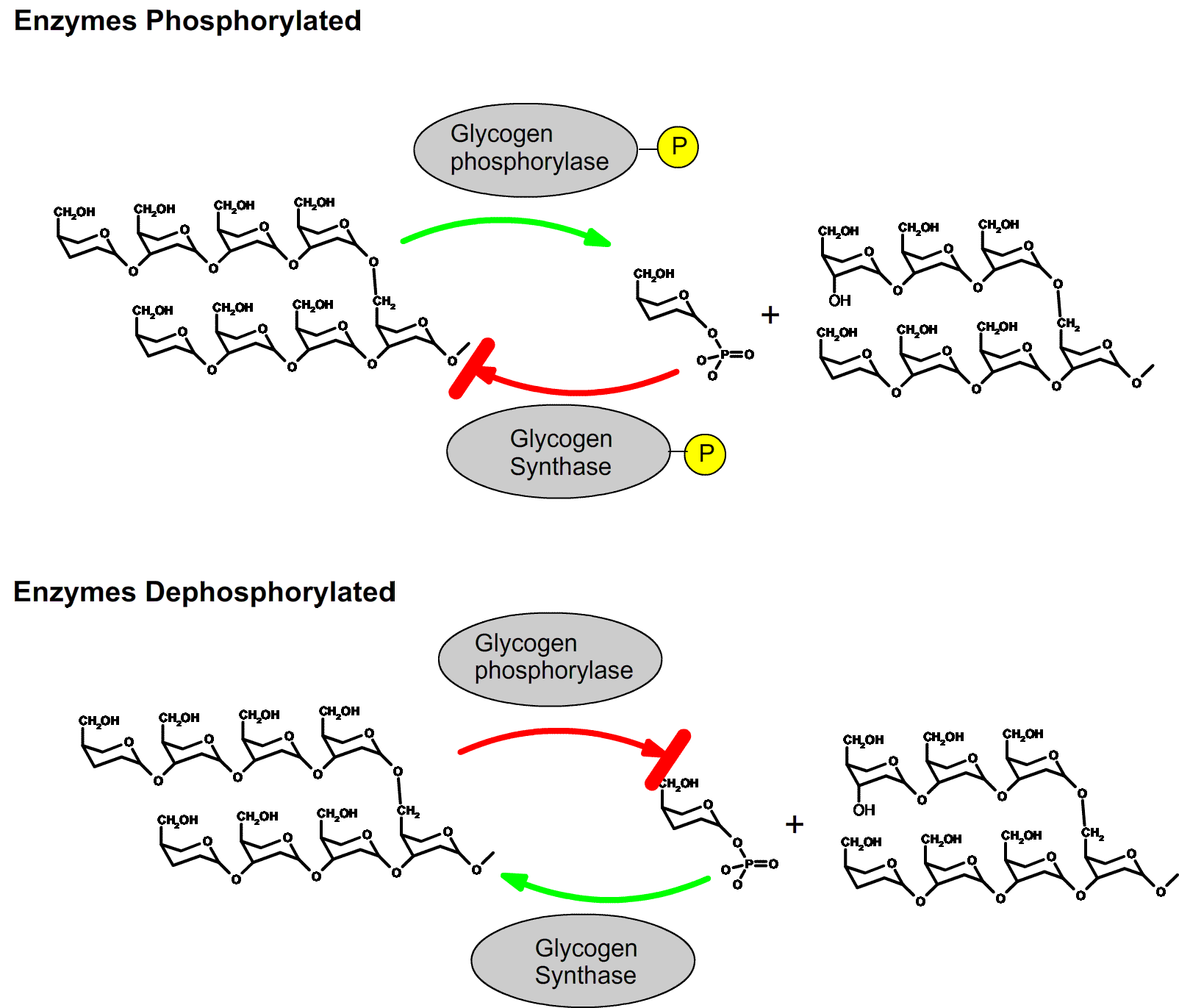 Glycogen regulation by phosphorylation of enzymes. Glycogen phosphorylase is active
when phosphorylated. In contrast, the phosphorylation of glycogen synthase inactivates the
enzyme. Dephosphorylation leads to a reversal in the regulation; inactivation of glycogen
phosphorylase and activation of glycogen synthase.
Glycogen regulation by phosphorylation of enzymes. Glycogen phosphorylase is active
when phosphorylated. In contrast, the phosphorylation of glycogen synthase inactivates the
enzyme. Dephosphorylation leads to a reversal in the regulation; inactivation of glycogen
phosphorylase and activation of glycogen synthase.
Glucose Synthesis - Gluconeogenesis
During glycolysis glucose is oxidized to pyruvate. The reverse reaction, the synthesis
of glucose from pyruvate occurs in the gluconeogenesis pathway. The major source of
pyruvate in the liver is lactate that was produced by active muscle during anaerobic
metabolism.
Since the process of converting glucose to pyruvate via glycolysis is energy releasing,
the reverse reaction must require energy. Most of the steps in the gluconeogenesis pathway
have a Gibbs free energy of nearly zero, and are easily reversed by slight changes in the
concentrations of products and reactions. However, there are three steps in glycolysis
that occur with such a large decrease in energy that they cannot become spontaneous in the
reverse direction. These steps are:
- The formation of glucose-6-P from glucose, coupled to ATP hydrolysis.
- The formation of fructose-1-6-P from fructose-6-P, coupled to ATP hydrolysis.
- The formation of pyruvate from phosphoenolpyruvate (PEP), releasing sufficient energy
to produce ATP.
Since the first two reactions use ATP to drive an otherwise unfavorable reaction, the
phosphorylation of sugars, the simpler reverse reaction, hydrolysis, will be favorable and
spontaneous. Consequently, in gluconeogenesis these two steps are catalyzed by
phosphatases instead of kinases. The third reaction in glycolysis, phosphoenol pyruvate
(PEP) to pyruvate, has such a large decrease in energy that it cannot be easily reversed
and must be accomplished by a different method.
In gluconeogenesis the conversion of pyruvate to PEP in gluconeogenesis occurs by in two
steps:
- The enzyme pyruvate carboxylase catalyzes: pyruvate + HCO3- +
ATP → oxaloacetate + ADP + Pi
- The enzyme PEP carboxykinase catalyzes: oxaloacetate + GTP → phosphoenolpyruvate + GDP
+ CO2.
Therefore a total of two "high energy" phosphate bonds (ATP/GTP → ADP/GDP +Pi)
are required to reverse this step.
The complete pathways for glucolysis and gluconeogenesis are shown below:
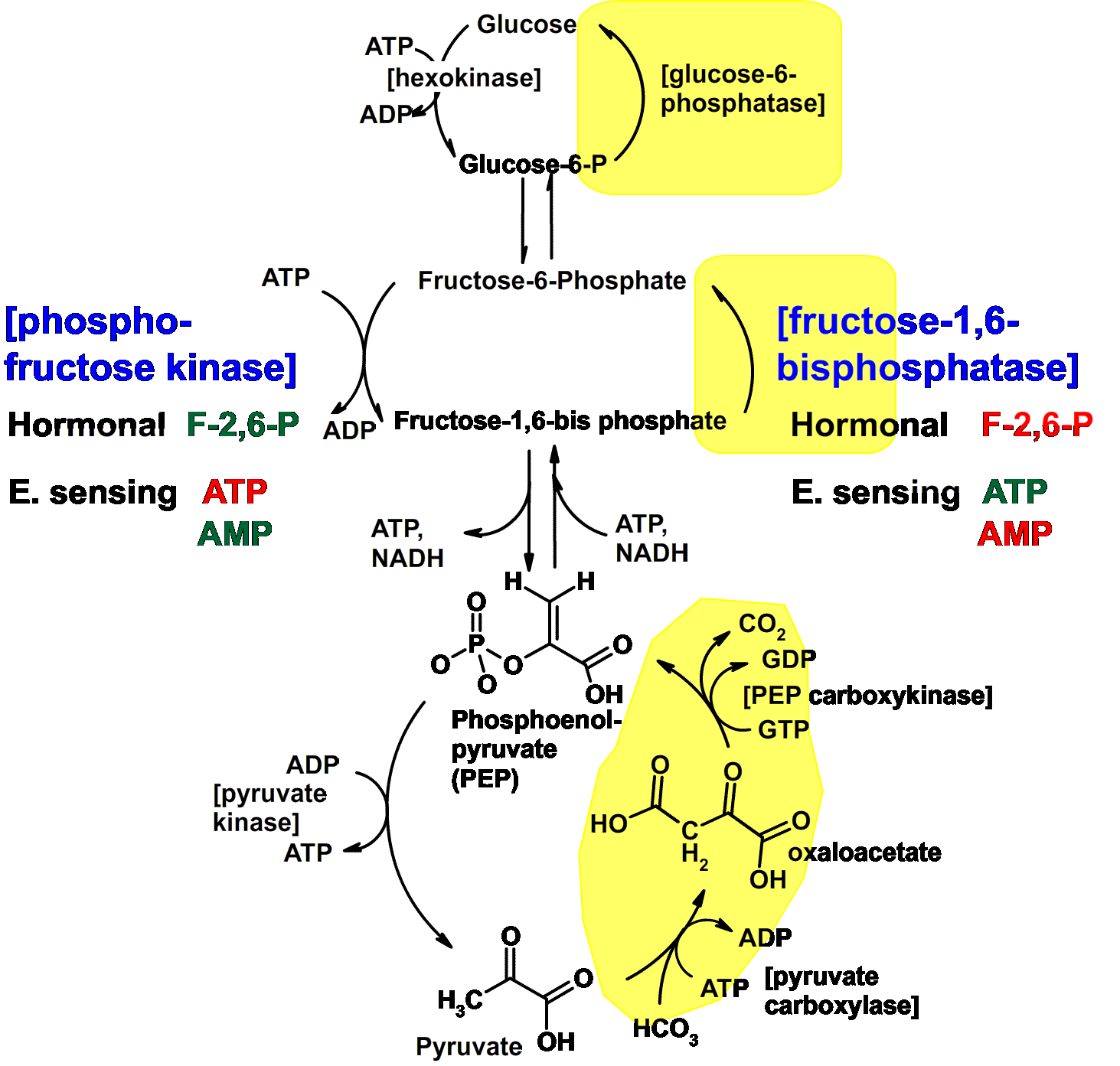 Glycolysis and Gluconeogenesis. The left part of the figure shows glycolysis, the
right side shows gluconeogenesis. Enzymatic steps that are accomplished by different
mechanisms (and enzymes) in each pathway are highlighted in yellow. The hydrolysis of
phosphate from fructose 1,6-P and glucose-6-P is spontaneous, consequently the kinases
used in glycolysis are replaced by phosphatases in gluconeogenesis. The conversion of
pyruvate to PEP follows a completely different route in gluconeogenesis than in
glycolysis. The enzymes whose names are colored red, phosphofructose kinase (PFK) and
fructose 1,6 bisphosphatase, catalyze important regulated steps in each
pathway.
Glycolysis and Gluconeogenesis. The left part of the figure shows glycolysis, the
right side shows gluconeogenesis. Enzymatic steps that are accomplished by different
mechanisms (and enzymes) in each pathway are highlighted in yellow. The hydrolysis of
phosphate from fructose 1,6-P and glucose-6-P is spontaneous, consequently the kinases
used in glycolysis are replaced by phosphatases in gluconeogenesis. The conversion of
pyruvate to PEP follows a completely different route in gluconeogenesis than in
glycolysis. The enzymes whose names are colored red, phosphofructose kinase (PFK) and
fructose 1,6 bisphosphatase, catalyze important regulated steps in each
pathway.
Glycolysis and gluconeogenesis are regulated at two levels.
First, at the cellular level, the regulatory enzymes are sensitive to
the energy levels of the cell. Second, at the organ level, the enzymes are regulated by hormones.
However, unlike glycogen metabolism, this regulation is not via
protein phosphorylation but by changes in levels of the allosteric
regulatory compound fructose 2,6 bisphosphate (F26P). The levels of
F26P are indirectly regulated by hormones by virtue of the fact that the
activity of the enzymes that make and degrade F26P are a affected by
protein phosphorylation.
Regulation by Energy Sensing. Cells have a relatively constant
amount of adenosine nucleotides, i.e. the sum of the concentrations of
AMP, ADP, and ATP are relatively constant. When the cell is consuming
energy, ATP is being hydrolyzed to ADP or to AMP. Consequently, high
levels of AMP and ADP indicate that the cell needs to activate
glycolysis to restore the levels of ATP. If ATP levels are high, then
glycolysis should be turned off. The excess ATP can be used to
synthesize glucose and other molecules if needed.
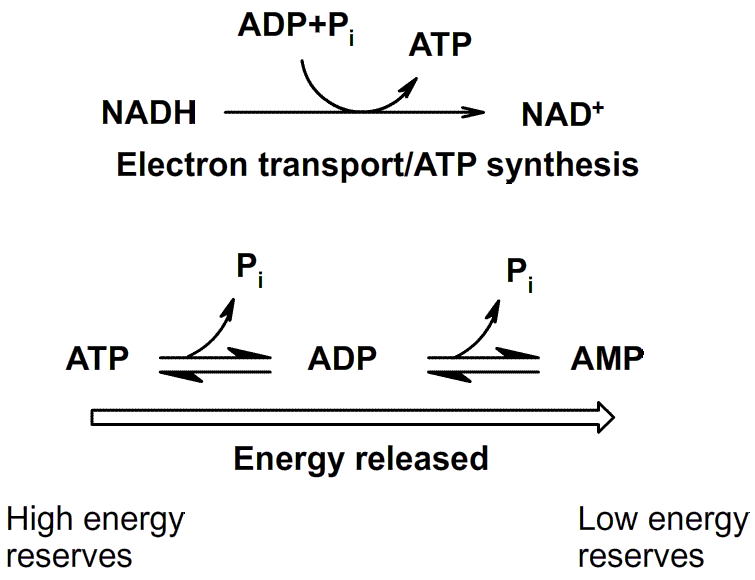 Cellular energy levels are high when ATP or NADH levels are high.
Cellular activities convert ATP to ADP and AMP. ATP levels can be
replenished by oxidation of NADH by electron transport and ATP
synthesis. When both NADH and ATP levels are low, the cell has low
energy reserves.
Cellular energy levels are high when ATP or NADH levels are high.
Cellular activities convert ATP to ADP and AMP. ATP levels can be
replenished by oxidation of NADH by electron transport and ATP
synthesis. When both NADH and ATP levels are low, the cell has low
energy reserves.
The enzymes phosphofructose kinase and fructose-1,6-bisphosphatase are under allosteric control by adenosine nucleotides:
High ATP
- Inhibits PFK - therefore glycolysis is off
- Activates fructose bisphosphatase - therefore gluconeogenesis can occur
HIgh AMP (low ATP)
- Activates PFK - glycolysis is turned on to regenerate ATP
- Inhibits fructose bis phosphatase - therefore gluconeogenesis is turned off.
Note that these two enzymes are regulated in a coordinated manner
such that only one pathway, glycolysis or gluconeogenesis, is turned on
at one time.
Regulation by Hormonal Control. The compound fructose -2,6
bisphosphate (F26P) is an allosteric activator of PFK. F26P also
inhibits fructose bisphosphatase, the key regulatory enzyme in
gluconeogenesis. The levels of F26P are controlled by hormones via
protein phosphorylation levels. When protein phosphorylation levels are
high, F26P is degraded, when phosphorylation levels are low, F26P is
synthesized.
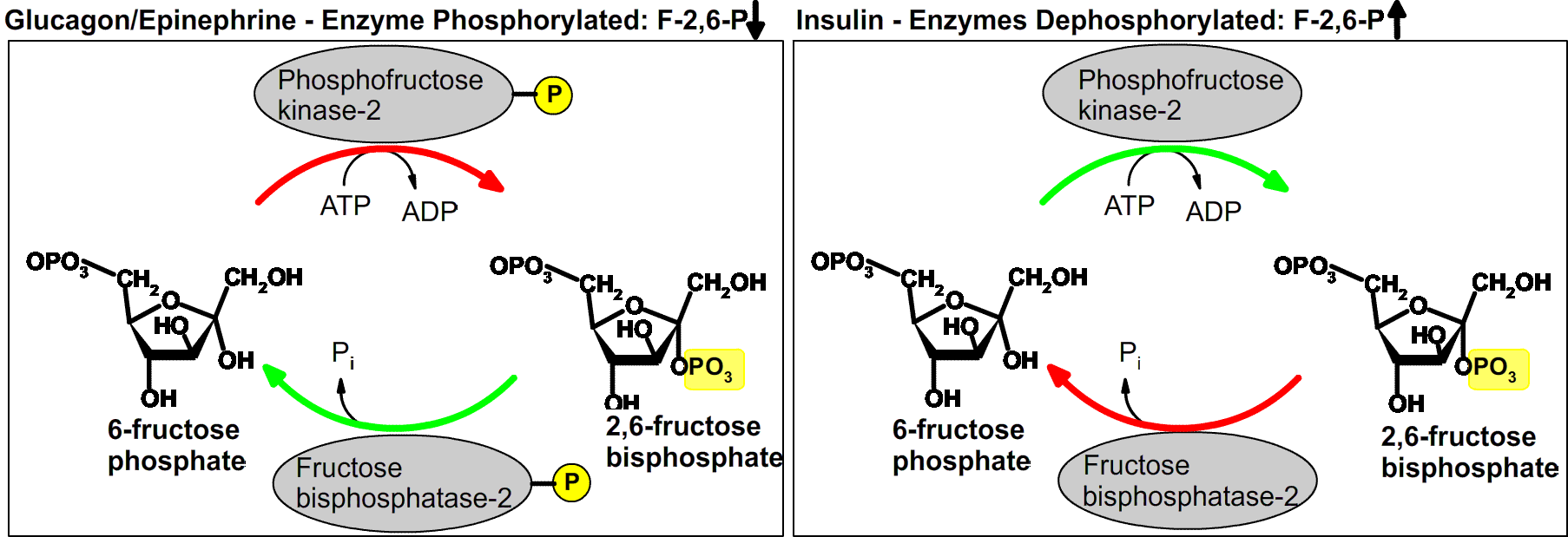 Regulation of fructose-2,6-bisphosphate (F26P) levels by hormones.
Glucagon and epinephrine cause phosphorylation of enzymes. The enzyme
that degrades (F26P) is activated when phosphorylated, consequently
F26P levels drop when either of these hormones are present. The reverse
occurs when insulin is present, the kinase becomes activated and F26P
levels rise. Note that F26P is generated from fructose-6-phosphate
(F6P), which is also an intermediate in glycolysis. Since the enzyme
that synthesizes F26P has the same function as PFK in glycolysis, adding
a phosphate to fructose-6-P, it is called PFK-2.
Regulation of fructose-2,6-bisphosphate (F26P) levels by hormones.
Glucagon and epinephrine cause phosphorylation of enzymes. The enzyme
that degrades (F26P) is activated when phosphorylated, consequently
F26P levels drop when either of these hormones are present. The reverse
occurs when insulin is present, the kinase becomes activated and F26P
levels rise. Note that F26P is generated from fructose-6-phosphate
(F6P), which is also an intermediate in glycolysis. Since the enzyme
that synthesizes F26P has the same function as PFK in glycolysis, adding
a phosphate to fructose-6-P, it is called PFK-2.
The regulation of glycolysis and gluconeogenesis by energy sensing and indirect hormonal control is summarize below.
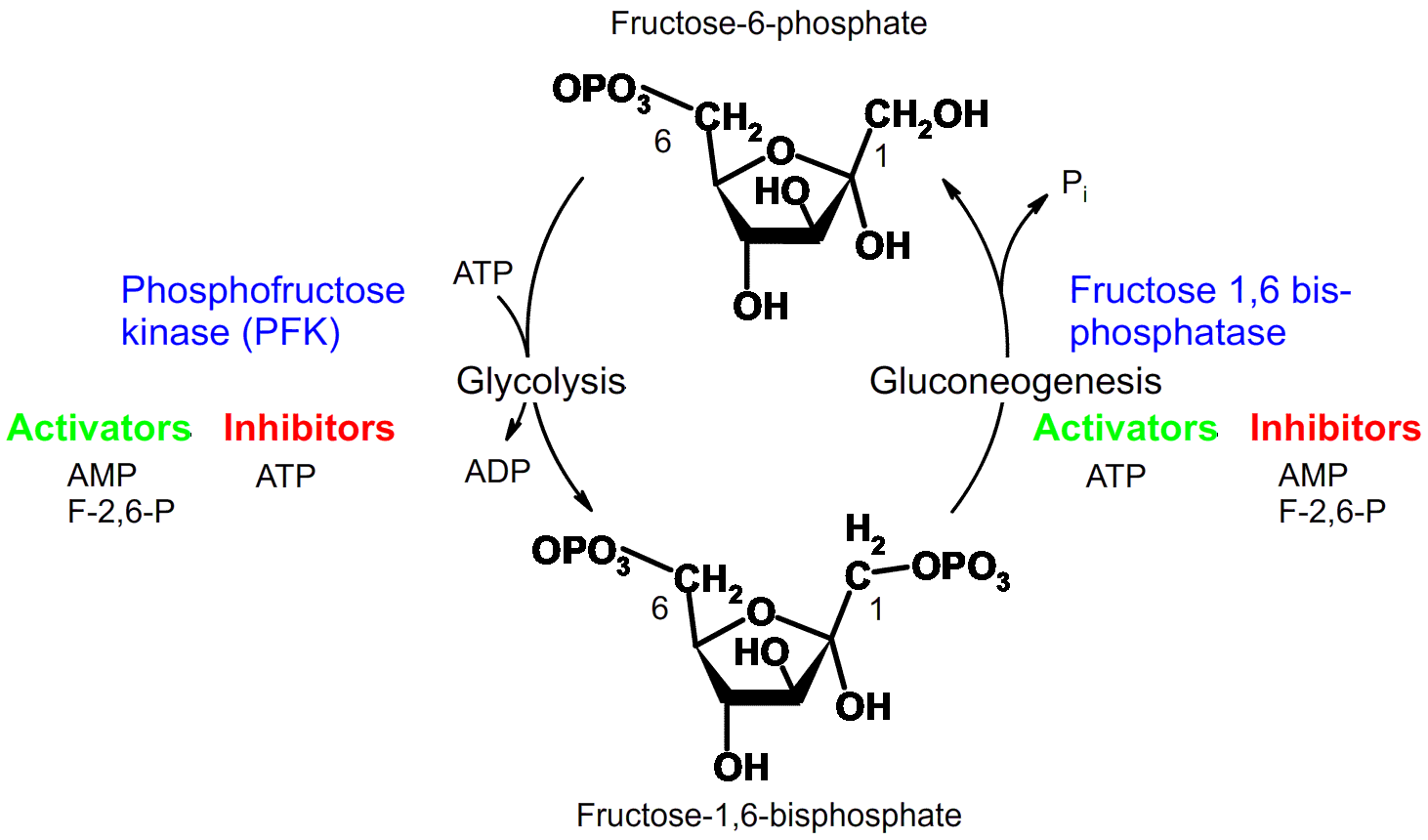 Regulation of PFK in glycolysis and fructose 1,6 phosphatase in
gluconeogenesis by energy sensing and fructose-2,6-bisphosphate levels.
Note that when one pathway is activated, the other is inhibited.
Regulation of PFK in glycolysis and fructose 1,6 phosphatase in
gluconeogenesis by energy sensing and fructose-2,6-bisphosphate levels.
Note that when one pathway is activated, the other is inhibited.
The coordinated regulation of glycolysis and gluconeogenesis dovetails
perfectly with the direct hormonal control of glycogen metabolism to
insure the liver cell responds appropriately to hormonal signals, i.e.
the production and release of glucose when glucagon or epinephrine are
present, or the storage and, if required, the oxidation of glucose when
insulin is present.
 The left panel shows the steps that occur when there is a demand for
glucose from the liver, as signaled by the hormones glucagon or
epinephrine. The end result is release of glucose from glycogen and the
activation of gluconeogenesis if there are sufficient ATP reserves.
The right panel shows the steps that occur when the liver (and muscle)
are to store excess glucose, as signaled by the hormone insulin. The
end result is the synthesis of glycogen and, if the cell requires energy
(ATP), the activation of glycolysis.
The left panel shows the steps that occur when there is a demand for
glucose from the liver, as signaled by the hormones glucagon or
epinephrine. The end result is release of glucose from glycogen and the
activation of gluconeogenesis if there are sufficient ATP reserves.
The right panel shows the steps that occur when the liver (and muscle)
are to store excess glucose, as signaled by the hormone insulin. The
end result is the synthesis of glycogen and, if the cell requires energy
(ATP), the activation of glycolysis.
Obesity and the Metabolism of Fats, Amino acids, and Complex Sugars
Fats are stored in cells as triglycerides, the condensation
product of three fatty acids with the three carbon polyalcohol
glycerol. Hormonal stimulation by glucagon or epinephrine activates
lipases which release the fatty acids from the glycerol. The fatty
acids are activated for oxidation in the cytosol by the addition of an coenzyme A group to the carboxylic acid, forming acyl-CoA.
After activation, the activated fatty acid is transported to the
mitochondria where it is subject to a pathway called beta-oxidation.
This process releases acetyl-CoA molecules from the fatty acid,
generating NADH and FADH2. The acetyl-CoA enters the TCA cycle for further oxidation.
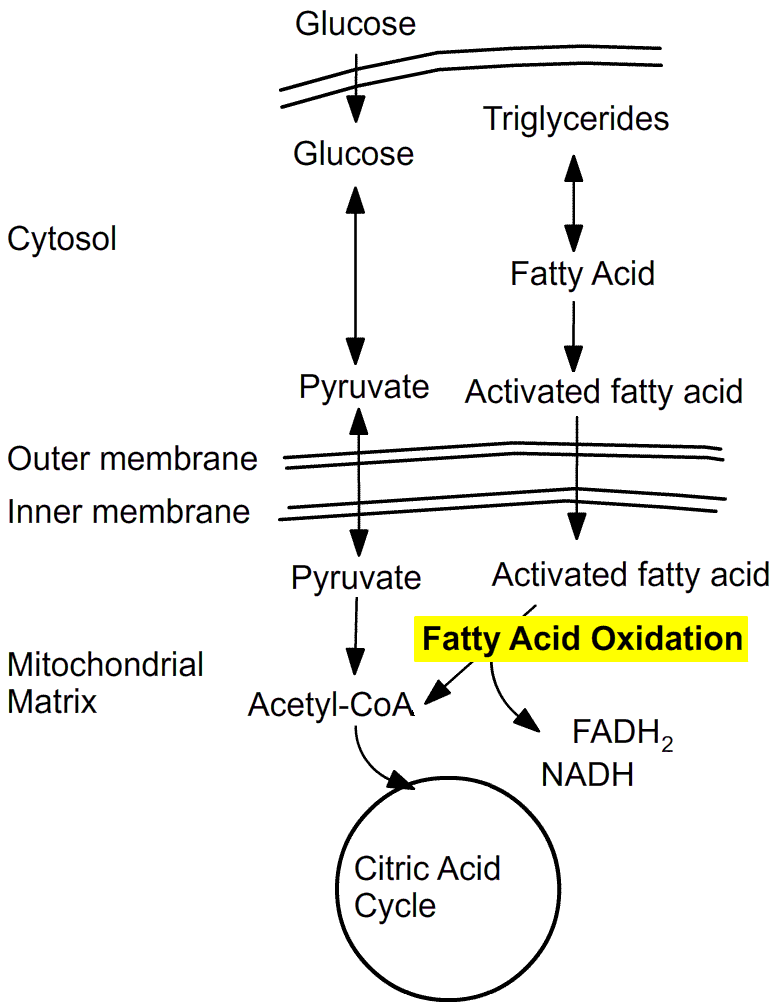 Fatty acids are stored in the cytosol as triglycerides. Fatty
acids are released from triglycerides by the action of lipases. To
begin the oxidation process, the fatty acid is activated by converting
the carboxylic acid to thioester to coenzymeA, generating acyl-CoA. The
acyl-CoA is transported into the mitochondrial matrix where oxidation
occurs.
Fatty acids are stored in the cytosol as triglycerides. Fatty
acids are released from triglycerides by the action of lipases. To
begin the oxidation process, the fatty acid is activated by converting
the carboxylic acid to thioester to coenzymeA, generating acyl-CoA. The
acyl-CoA is transported into the mitochondrial matrix where oxidation
occurs.
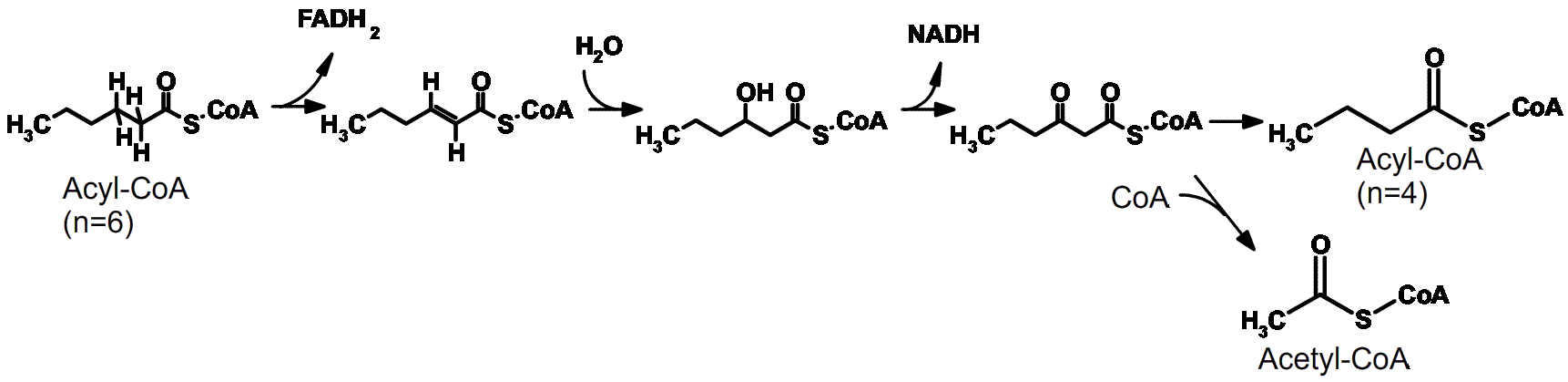 Fatty acids are stored in the cytosol as triglycerides. Fatty
acids are released from triglycerides by the action of lipases. To
begin the oxidation process, the fatty acid is activated by converting
the carboxylic acid to thioester to coenzymeA, generating acyl-CoA. The
acyl-CoA is transported into the mitochondrial matrix where oxidation
occurs.
Fatty acids are stored in the cytosol as triglycerides. Fatty
acids are released from triglycerides by the action of lipases. To
begin the oxidation process, the fatty acid is activated by converting
the carboxylic acid to thioester to coenzymeA, generating acyl-CoA. The
acyl-CoA is transported into the mitochondrial matrix where oxidation
occurs.
Fatty acids can also be synthesized from acetyl-CoA by an energy
requiring process. As with glycolysis/gluconeogenesis many of the same
enzymes are utilized in both pathways. However, highly exergonic
reactions in the degradative pathway are accomplished by a different
mechanism in the synthetic pathway.
Complex carbohydrates are converted to monosaccharides
and enter glycolysis at a number of different points. Most
monosaccharides enter glycolysis above the step catalyzed by
phosophofructose kinase (PFK). Consequently, the degradation of
practically all sugars is controlled by this key regulatory enzyme.
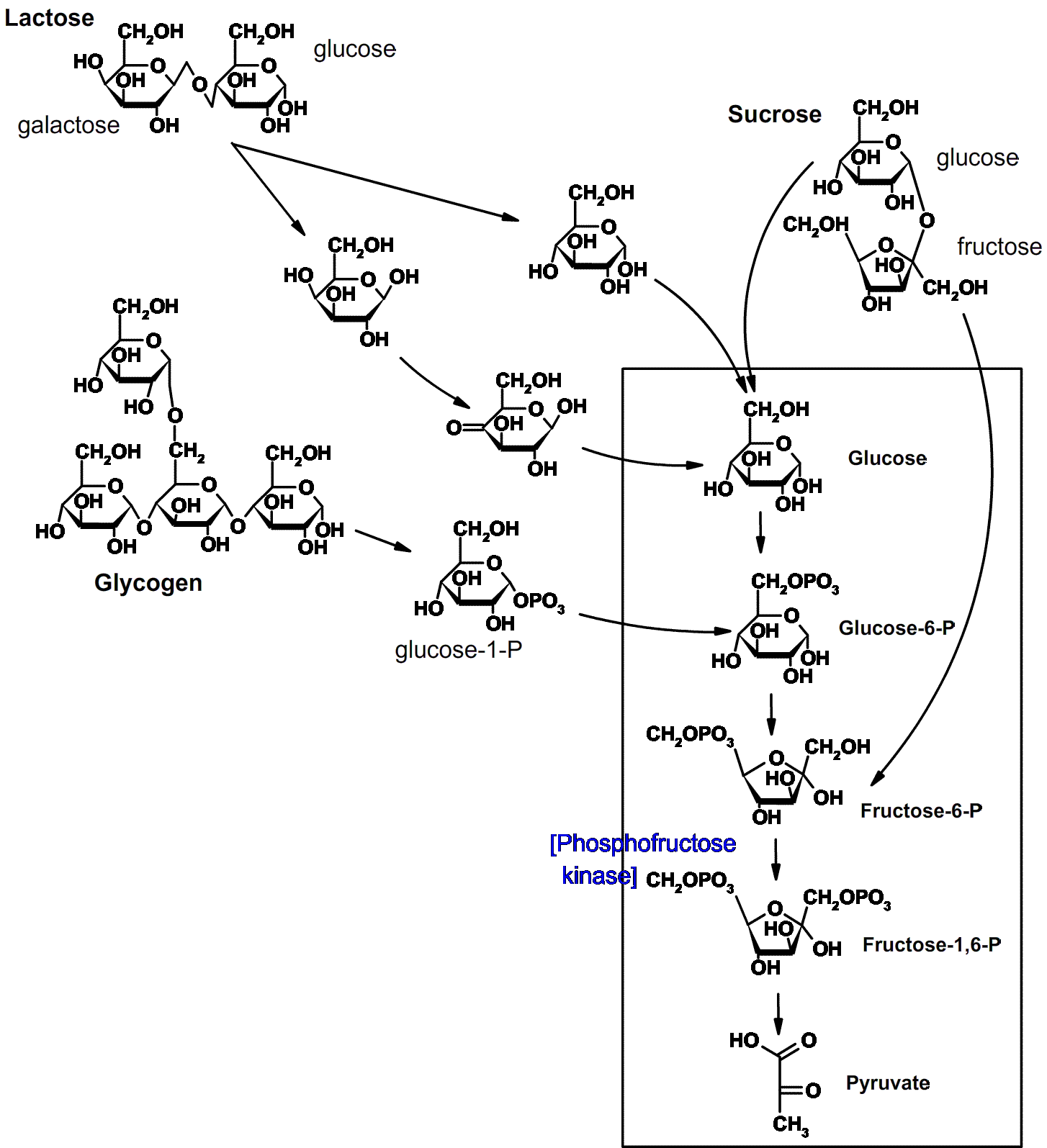 Metabolism of complex carbohydrates. Glycogen releases glucose
as glucose-1-P which can be readily converted to glucose-6-P and enter
glycolysis. Sucrose, or table sugar, is broken down into glucose and
fructose. The fructose is phosphorylated and enters glycolysis above
the PFK step. Lactose, the major sugar in milk, provides one glucose
that enters glycolysis directly and one galactose. The chiral center at
the fourth carbon of galactose is then inverted, creating glucose.
Metabolism of complex carbohydrates. Glycogen releases glucose
as glucose-1-P which can be readily converted to glucose-6-P and enter
glycolysis. Sucrose, or table sugar, is broken down into glucose and
fructose. The fructose is phosphorylated and enters glycolysis above
the PFK step. Lactose, the major sugar in milk, provides one glucose
that enters glycolysis directly and one galactose. The chiral center at
the fourth carbon of galactose is then inverted, creating glucose.
Although the metabolism of most amino acids is a
complicated multi-step process, a number of amino acids enter the main
oxidative pathways by a single transaminase reaction. In the transaminase reaction
a ketone group is replaced by an amino group. For example, the amino
acid alanine is converted to pyruvate by the following transaminase
reaction:
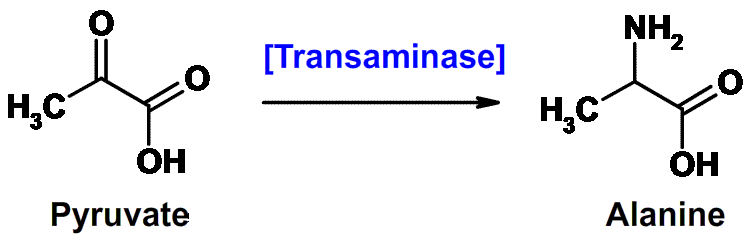 Alanine can be reversibly converted to pyruvate by transamination.
Alanine can be reversibly converted to pyruvate by transamination.
Why do Humans Get Fat?
A key feature of human metabolism is the fact that the conversion of pyruvate to acetyl-CoA is irreversible.
Consequently, once ingested carbon from any source, fats, protein, or
carbohydrates, is converted to acetyl-CoA it has only two fates:
- Entry into the TCA cycle for oxidation and the production of energy.
- Storage as triglycerides, in specialized adipose tissue.
The TCA cycle is regulated by energy sensing. When the levels of the
high energy compounds ATP or NADH become elevated the TCA cycle halts -
there is no need for the cell to produce energy. Under these
conditions, any excess acetyl-CoA is stored as fat, regardless of the
source of the acetyl-CoA. The ingestion of excess amounts of proteins
or carbohydrates result in the generation of fat.
Fats are a less desirable form of energy storage since the
production of energy from fats is slower than the production from
glucose. In addition, a number of tissues in the body, in particular
the brain, use glucose as their principal energy source. Consequently,
the enzyme that converts pyruvate to acetyl-CoA is tightly regulated.
Pyruvate dehydrogenase is inhibited by:
- Product inhibition by NADH and acetyl-CoA.
- Phosphorylation of the enzyme causes inactivation.
Consequently, pyruvate will not be converted to acetyl-CoA if there
is excess NADH or acetyl-CoA. Furthermore, the hormones glucagon and
epinephrine, which evoke protein phosphorlation, prevent the loss of
pyruvate to acetyl-CoA such that pyruvate can be used for the synthesis
of glucose.
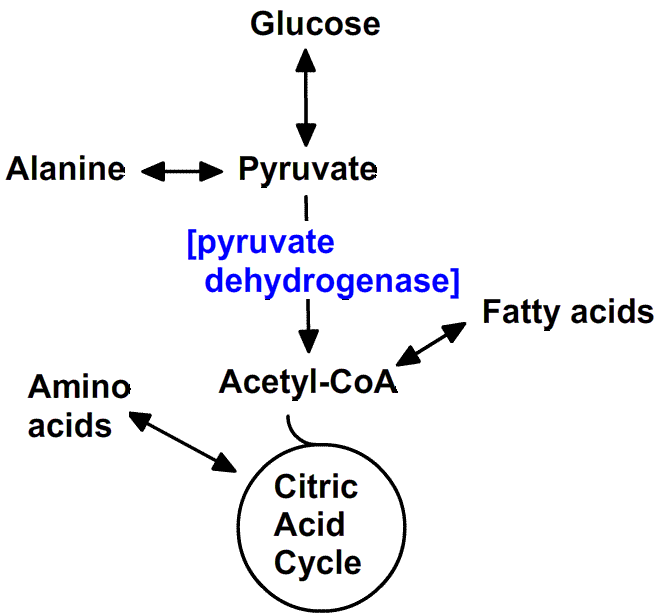 Fats are synthesized from excess amino acids and sugars.
Fats are synthesized from excess amino acids and sugars.
Although pyruvate dehydrogenase is well regulated, it is clear by
observation of the human population that the flow of carbon from
pyruvate to acetyl-CoA to fatty acids still occurs.
This page is dedicated to instructions about the Activities.
Make sure your browser supports MathML
Many pages of this course contain formulas which require that your
browser supports displaying MathML. You should run "Test and Configure
My System" from your My Courses page to make sure your browser supports
the tools the course uses.
Types of Activities
This course has many interactive activities that you are encouraged to complete.
You can repeat many of the activities as necessary. General instructions for the
use of the tools will always be available in the Glossary Unit: Instructions for
Activities.
- "Learn By Doing" and "Explore further" These links take you to exercises
you can repeat multiple times. They will usually be interactive and help
you to understand the concept being discussed. These activities are "low
stakes." You will not be graded on these activities.
- "Did I Get This" links take you to self-assessment questions. These
exercises are graded for your information only; the grade is not
recorded in the grade book. Many of the questions will contain hints and
feedback to help you guide your learning and review of the material. You
may take these quizzes as many time as you like.
- "Many Students Wonder" are activities that provide additional detail or
material. These materials supplement the ideas being presented with
articles showcasing current research, items in the news, or other
interesting tidbits.
- "Quiz" This is a graded assessment. The grade will be recorded in the
gradebook, and you can take these quizzes only one time. You should take
these quizzes only after you have mastered the other activities such as
the "Learn By Doing" and "Did I Get This." These assessments are not
available in the Open and Free version of the course.
- "My Response" This activity serves two functions. The My Response
buttons in the content follow questions posed in the text. You should
treat these questions as practice for the exam. Your first response will
be saved for you to view in the future. The responses are not graded but
are visible to the instructors and may be used in classroom dicussion.
The second use of the "My Response" button is at the end of a lesson
just before a graded exam. This activity is you chance to send questions
to the instructors. The questions can be for clarification of the
material or for something you find confusing. The questions can also be
for further information. The questions you pose are visible to the
instructors may be used for in class discussion.
Flash Interaction
Any Flash movie can be zoomed.
- Mac users: "Control" click on the Flash movie and select the "Zoom In"
option in the menu.
- PC users: Right-click on the Flash movie for the menu of options.
Jmol 3-D Molecule Viewer
Jmol is an open-source Java viewer for chemical structures in 3D. Jmol returns a
3D representation of a molecule that may be manipulated in a number of ways.
- Mac users: Click and drag to rotate the molecule. Option drag to zoom in or
out. "Control" click on the Jmol for the menu of options.
- PC users: Click and drag to rotate the molecule. Alt drag to zoom in or out.
Right-click on the Jmol for the menu of options.
FlashMol 3-D Molecule Viewer
FlashMol allows you to see chemical structures in 3-D and to rotate the structure
in all three dimensions.
Click an hold an area of
the 3-D molecule and drag the mouse cursor in the direction you want the
molecule to move. Compare the 3-D molecule to the text representation.
In the Functional
Groups Activity you can find many examples of 2-D and 3-D
representations.
The following is the list of terms defined in this course.
- Aldose
-
(Definition)
A category of simple carbohydrates where the number 1 carbon (the top carbon)
contains the carbonyl that is flanked by a hydrogen and a carbon thus making
this an aldehyde.
Refer to
page.
- Allosteric Binding
-
(Definition)
Allosteric binding causes conformational changes in an enzyme that can either
inhibit or activate the enzyme.
Refer to page.
- Catalyst
-
(Definition)
A catalyst is a participant in a chemical reaction that speeds up the
reaction but is not consumed itself. Enzymes are biological catalysts that
mediate the conversion of substrate to product, by lowering the activation
energy of the reaction. Enzymes re not consumed or modified during the
process. Enzymes are generally made of proteins but nucleic acid enzymes,
ribozymes, have also been discovered.
Refer to page.
- Chirality
-
(Definition)
When identical groups attached to a carbon are arranged in multiple ways such
that two of the resulting structures are non-superimposable, they are mirror
images of each other.
Refer to page.
- Covalent bond
-
(Definition)
Covalent bonds represent the sharing of the electrons (negatively charged
subatomic particles between atoms.) The number of covalent bonds that can
form is dictated by the number of unpaired electrons in the outer valence
shell of the atom.
Refer to page.
- De novo synthesis
-
(Definition)
De novo is new synthesis from simple molecules such as deoxynucleotides in this
case without a starting place - the primer.
Refer to page.
- Electronegativity
-
(Definition)
The tendency of an atom to attract electrons to itself. Electronegativity
increases as one moves from left to right, across the periodic chart.
Because the electronegative atoms have the potential to attract electrons,
i.e., the electrons spend more time on the electronegative atom, a molecule
containing an electronegative atom will have partial negative charge
associated with that atom (as indicated by δ
-). The the
bonding partner in the covalent bond becomes partially positively charged
(as indicated by δ
+ ).
Refer to page.
- Eukaryotes
-
(Definition)
An organism with complex cells with distinctive traits such as a nucleaus,
membrane-bound organelles, a cytoskeleton, and the presence of introns in genes.
Refer to
page.
- Furanose
-
(Definition)
Cyclic sugars that contain a five membered ring are called furanoses.
Refer to
page.
- Genome
-
(Definition)
the complete DNA sequence of one set of chromosomes from an organism.
Refer to
page.
- Hydrogen Bond
-
(Definition)
The attraction of an electronegative atom for a hydrogen that is covalently
bonded to another electronegative atom. This involves the attraction of a
hydrogen with a partial positive charge to an atom with a partial negative
charge. However, only hydrogens covalently bonded to an electronegative atom
can participate in hydrogen bonding.
Refer to page.
- Hydrophilic
-
(Definition)
We need a good definition and image here Because polar molecules are
generally water soluble, they are referred to as being hydrophilic, or
water-loving.
Refer to
page.
- Hydrophobic
-
(Definition)
Hydrophobic, or water-fearing, molecules do not interact with water and are
characterized by a complete lack of electronegative atoms. In aqueous
solutions the hydrophobic molecules are driven together to the exclusion of
water.
Refer to
page.
- Ionic Bond
-
(Definition)
An
ion is an atom or molecule which has lost or gained one or more
electrons, making it positively or negatively charged.
We need a complete
definition
Ionic bonds form between oppositely charged atoms. No electron sharing or
transfer occurs. The atoms are attracted to each other due to their opposite
charges.
Refer to
page.
- Ketose
-
(Definition)
A carbohydrate with a carbonyl at the number 2 carbon (the center carbon)
that is flanked by carbons on both sides, thus making this carbonyl a
ketone, is a ketose. This is a polar, hydrophilic, water-soluble molecule.
Refer to
page.
- Polymer
-
(Definition)
These are large molecules consisting of repeating structural units, or
monomers, connected by covalent chemical bonds.
Refer to page.
- Prokaryotes
-
(Definition)
A unicellular organizm lacking certain complex cell features, such as a
membrane-bound nucleus and gene introns.
Refer to page.
Biologists have different ways of representing chemical structures. Each type conveys
different information.
Use this as a reference for the important functional groups
and representative molecules for each group. You can select individual
groups or select by property. Inside each functional group there are
multiple examples of the group in actual molecules. The molecules are
represented as text and in 3-D.
The following is the list of tutorial animations explaining various complex biological
processes.
|
Biological Membranes Biological
membranes are dynamic structures composed of a diverse set of phospholipid
molecules and proteins. This tutorial explores some of the properties of the
membranes. Refer to
page.
|
|
|
Phase Transition
Phospholipids bilayers undergo a cooperative phase transition or melting that is
similar to protein denaturation. The high degree of cooperativity is due to
extensive interactions between the non-polar acyl chains in the center of the
bilayer. The overall structure of the lipid bilayer is not changed by this
transition. However, the disorder of the non-polar hydrocarbon chains increases
dramatically after melting. Refer to page.
|
|
|
Glucose Transport
The tutorial shows how the glucose transporter molecule is structured with a
spiral channel that allows the glucose molecules to passively navigate the
channel and move through the membrane. Refer to page.
|
|
|
Signal Transduction
This animation illustrates how binding of a ligand to its receptor ultimately
leads to the production of high levels of cAMP, an important second messenger in
the signal transduction pathway. Refer to transport page.
[ Refer to
signaling in metabolism page ]
|
|
|
Lactose Permease
Transporter The Escherichia coli lactose permease is an example
of secondary active transport (Campbell, p. 210). This enzyme is similar in
structure to others in the major facilitator superfamily (MFS) of transporters.
More than 1000 examples of MFS transporters have been identified in the genomes
of bacteria, plants, and animals. Refer to page.
|
|
|
Receptor Mediated
Endocytosis Many macromolecules are taken into the cell by a
process described as endocytosis. The macromolecules do not pass through the
membrane directly to the cytoplasm but instead are taken up by the cell,
processed by cell and then delivered to the cytoplasm. For many of the endocytic
processes, the uptake of the molecule is very specific and is controlled by
recognition of the molecule by a specific receptor on the surface of the cell.
This process is called Receptor Mediated Endocytosis. Refer to page.
|
|
|
ADP/ATP Exchange This
tutorial explains ADP/ATP exchange, a form of Antiport transport, a mechanism
where two different molecules facilitate each others pass through the membrane
in opposite directions. Refer to
page.
|
|
|
DNA Replication The
enzymatic and structural features of DNA replication in all organisms are very
similar. These animations illustrate the processes involved in the replication
of the Eschericia coli chromosome. The names of the DNA sites, enzymes, and
other protein factors differ in viruses and in eukaryotic organisms, however,
the basic features have been highly conserved in evolution. In E. coli, DNA
replication begins at a single site on the chromosome called "OriC".
Refer to
page.
|
|
|
DNA Transcription This
tutorial shows the steps involved in transcribing DNA into mRNA. Transcription
is illustrated using the E. coli lactose operon (lac) where transcription is
regulated negatively by the lac repressor and positively by CAP-cAMP complex.
Refer to
page.
|
|
|
RNA Translation - Protein
Synthesis The biosynthesis of proteins involves translation of
the information contained in the mRNA into a polypeptide sequence. Each triplet
of bases in the mRNA encodes one amino acid. The translation is accomplished by
the ribosome with the use of specialized RNA molecules called transfer RNA, or
tRNA. The tRNA molecules bind to the mRNA and deliver the correct amino acid to
the growing polypeptide chain. Refer to page.
|
|
|
ATP Synthesis The
synthesis of ATP utilizing the proton gradient as a source of energy is an
example of direct coupling. The energy released as the protons flow through the
enzyme cause a conformational change in the protein that causes the formation of
ATP from bound ADP and inorganic phosphate. Refer to page.
|
|
|
Serine Protease The
tutorial shows the chemical mechanism of serine proteases, enzymes that in the
family differ only in their substrate specificity.The Trypsin for example is an
extracellular protease that hydrolyzes peptide bonds during digestion in the
small intestine. |
|
|
Metabolism Overview The four key
pathways, glycolysis, the TCA cycle, electron transport, and ATP synthesis are
outlined in yellow. Glucose that is brought into the cell via the glucose
transporter can suffer two fates, oxidation or storage as glycogen. Oxidation
occurs in glycolysis and the TCA cycle, releasing the carbon atoms in glucose as
CO2. Note that oxygen is not used until the end of the electron transport chain.
High energy electrons, symbolized as orange balls are carried on organic
electron carriers to the electron transport chain. As these electrons move
through the four complexes, protons are pumped from the mitochondrial matrix
across the inner mitochondrial membrane. As these protons flow back through the
membrane via ATP synthase, ATP is generated. Refer to page.
|
|
The following is the list of simulations provided within the various modules of the
course.
|
Dissociation of Weak
Electrolytes Some molecules are weak electrolytes and exist in a
reversible equilibrium between the starting molecule and its dissociated parts.
For molecules that are weak electrolytes and act as acids (proton donors), the
ratio of the products of the dissociated parts and the parent molecule is a
constant (K) in neutral water for each separate molecular structure.
Refer to page.
|
|
|
Equilibrium of MO complex
formation Equilibrium state as determined by protein and ligand
concentrations: This simulation allows you to explore the equilibrium between
free protein, ligand, and the complex, and how ligand concentration affects the
equilibrium. Refer to page.
|
|
|
Enzyme Catalysis
Enzymes bind to substrates in a manner similar to the way myoglobin binds oxygen
or the estradiol binds to the estrogen receptor, but enzymes can go one step
further. In this case the ligand is specifically referred to as the substrate
(the molecule that the enzyme will convert to product) and it binds to a
specific binding region of the enzyme referred to as the active site. Once
bound, the ligand, or substrate, can either simply reversibly come off the
enzyme, or it can be converted into a new compound or product. Refer to page.
|
|
|
Membrane
Permeability The introduction of a cell or liposome to the
solution places a barrier to the molecules. As three different molecules diffuse
to equilibrium in the following simulation, they encounter the lipid bilayer
depicted by the horizontal membrane across the center of the stage. Note that
one type of molecule passes freely through the lipid bilayer while the second
type of molecule only occasionally passes through the membrane and the lipid
bilayer is totally impermeable to the third type of molecule. These molecules
move through the membranes via passive diffusion. Refer to page.
|
|
|
Osmosis: Isotonic
Equilibrium Cells continually encounter changes in their
external ionic environment and will spontaneously respond by attempting to
equalize the concentration of ions on the inside and outside of the cell.
Because the plasma membrane (lipid bilayer) is significantly less permeable to
ions than water, the establishment of an equal concentration of the ions on
either side of the membrane is accomplished by the net movement of water toward
the higher concentration of ions to reduce the concentration. This movement of
water in response to an imbalance of solute (ion) is referred to as osmosis.
Refer to page.
|
|
|
Intercellular Transport: Gap
Junctions This simulation demonstrates how transmembrane protein
structures from adjacent cells line up to form Gap Junctions, channels between
cells that act as size exclusion transporters. While water molecules are small
enough to move through the membranes, the Gap Junctions facilitate that movement
and the movement of molecules up to 1500 daltons (approximately a 15 amino acid
peptide) but not larger molecules. Refer to page.
|
|
|
Product inhibition This
simulation demonstrates how a compound that is further down the pathway, or even
a compound in a separate pathway, can inhibit a reaction Refer to page.
|
|
|
Allosteric Binding This
simulation demonstrates how Allosteric binding causes conformational changes to
an enzyme that can inhibit or activate Enzyme-substrate binding. Refer to page.
|
|
|
Calculating Free Energy This
simulation demonstrates how to calculate the Gibbs free energy in a controlled reaction. Refer to page.
|
|
Use this table as a quick reference for the major elements.
| Atomic Properties of the major biological atoms |
|---|
| Atom |
Mass |
Covalent Bonds |
Electronegativity |
| H |
1 |
1 |
2.20 |
| C |
12 |
4 |
2.55 |
| N |
14 |
3 or 4 (e.g. NH4+) |
3.04 |
| O |
16 |
2 |
3.44 |
| P |
31 |
5 |
2.10 |
| S |
32 |
2 |
2.58 |
|
 This is a mouse tissue culture cell showing the cytoskeletal structure. The
large clearing in the center is the nucleus. Image courtesy of Jon Jarvik,
Department of Biological Sciences, Carnegie Mellon University.
This is a mouse tissue culture cell showing the cytoskeletal structure. The
large clearing in the center is the nucleus. Image courtesy of Jon Jarvik,
Department of Biological Sciences, Carnegie Mellon University.